Crystal structures of Arabidopsis and Physcomitrella CR4 reveal the molecular architecture of CRINKLY4 receptor kinases.
Okuda, S., Hothorn, L.A., Hothorn, M.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Predicted protein | 383 | Physcomitrium patens | Mutation(s): 0 Gene Names: PHYPA_009221, PHYPADRAFT_159650 EC: 2.7.11.1 | 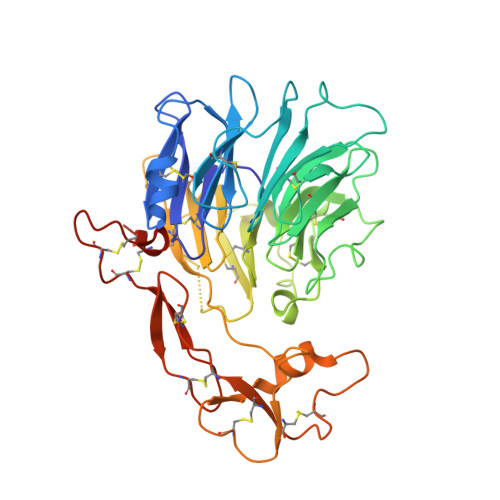 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A9RKG8 | ||||
Glycosylation | |||||
| Glycosylation Sites: 3 | |||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| alpha-L-fucopyranose-(1-6)-2-acetamido-2-deoxy-beta-D-glucopyranose | I, K, M, O, Q I, K, M, O, Q, S, T, U | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G86851RC GlyCosmos: G86851RC GlyGen: G86851RC | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Length | 2D Diagram | Glycosylation | D Interactions |
| 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose | J, N, P, R | 2 |  | N-Glycosylation | |
Glycosylation Resources | |||||
GlyTouCan: G42666HT GlyCosmos: G42666HT GlyGen: G42666HT | |||||
| Ligands 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | CA [auth B] EA [auth C] GA [auth D] HA [auth E] IA [auth F] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| MLT Download:Ideal Coordinates CCD File | V [auth A] | D-MALATE C4 H6 O5 BJEPYKJPYRNKOW-UWTATZPHSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | AA [auth B] BA [auth B] DA [auth B] FA [auth C] LA [auth G] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 88.579 | α = 90 |
| b = 183.962 | β = 96.05 |
| c = 98.173 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Swiss National Science Foundation | Switzerland | 31003A_176237 |
| Swiss National Science Foundation | Switzerland | 31CP30_180213 |