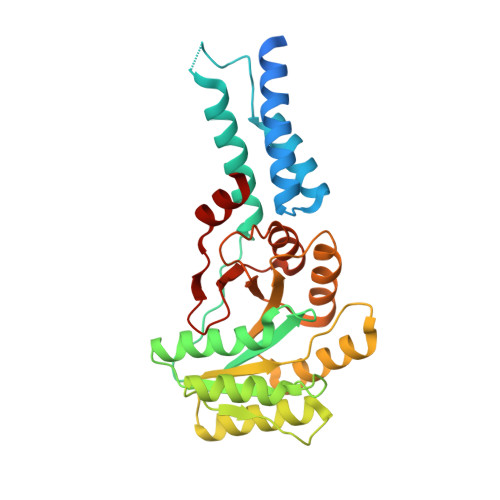
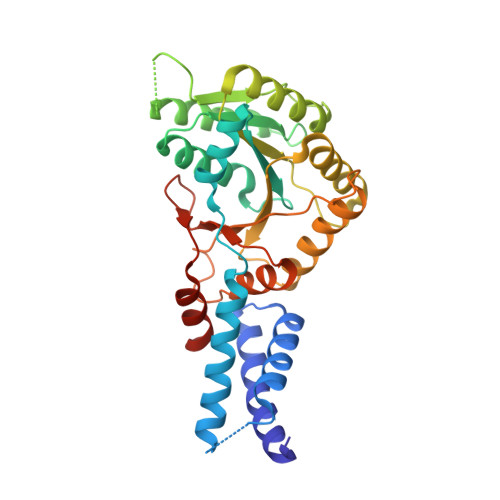
Structural and Functional Impact of SRP54 Mutations Causing Severe Congenital Neutropenia.
Juaire, K.D., Lapouge, K., Becker, M.M.M., Kotova, I., Michelhans, M., Carapito, R., Wild, K., Bahram, S., Sinning, I.(2021) Structure 29: 15
- PubMed: 33053321 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2020.09.008
- Primary Citation Related Structures:
6Y2Z, 6Y30, 6Y31, 6Y32 - PubMed Abstract:
The SRP54 GTPase is a key component of co-translational protein targeting by the signal recognition particle (SRP). Point mutations in SRP54 have been recently shown to lead to a form of severe congenital neutropenia displaying symptoms overlapping with those of Shwachman-Diamond syndrome. The phenotype includes severe neutropenia, exocrine pancreatic deficiency, and neurodevelopmental as well as skeletal disorders. Using a combination of X-ray crystallography, hydrogen-deuterium exchange coupled to mass spectrometry and complementary biochemical and biophysical methods, we reveal extensive structural defects in three disease-causing SRP54 variants resulting in critical protein destabilization. GTP binding is mostly abolished as a consequence of an altered GTPase core. The mutations located in conserved sequence fingerprints of SRP54 eliminate targeting complex formation with the SRP receptor as demonstrated in yeast and human cells. These specific defects critically influence the entire SRP pathway, thereby causing this life-threatening disease.
- Heidelberg University Biochemistry Center (BZH), Im Neuenheimer Feld 328, D-69120 Heidelberg, Germany.
Organizational Affiliation: