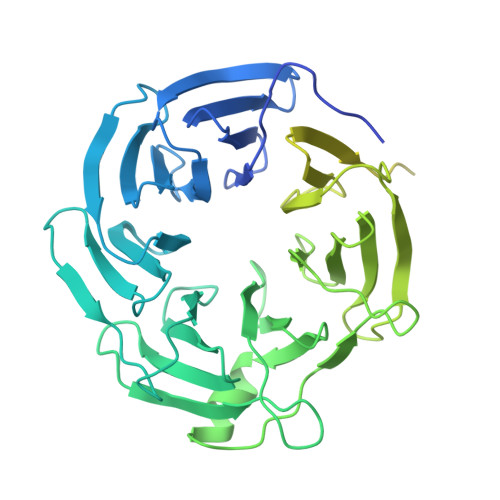
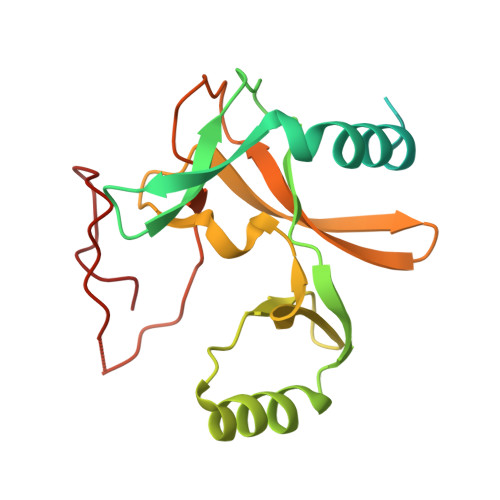
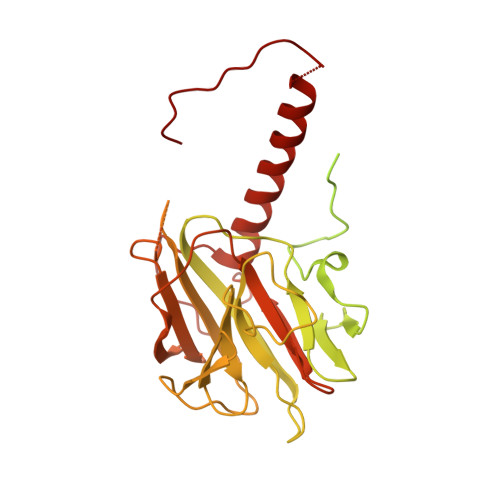
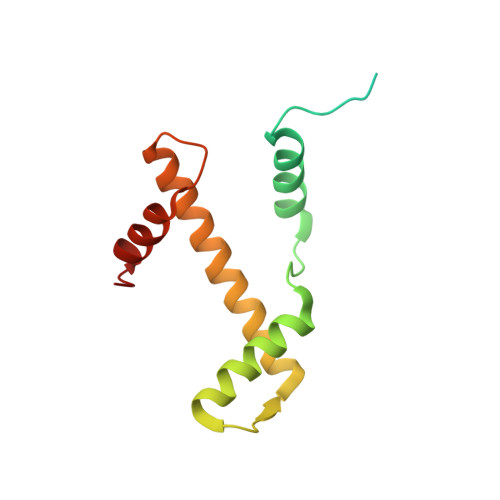
Mechanism for DPY30 and ASH2L intrinsically disordered regions to modulate the MLL/SET1 activity on chromatin.
Lee, Y.T., Ayoub, A., Park, S.H., Sha, L., Xu, J., Mao, F., Zheng, W., Zhang, Y., Cho, U.S., Dou, Y.(2021) Nat Commun 12: 2953-2953
- PubMed: 34012049 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-23268-9
- Primary Citation Related Structures:
6W5I, 6W5M, 6W5N - PubMed Abstract:
Recent cryo-EM structures show the highly dynamic nature of the MLL1-NCP (nucleosome core particle) interaction. Functional implication and regulation of such dynamics remain unclear. Here we show that DPY30 and the intrinsically disordered regions (IDRs) of ASH2L work together in restricting the rotational dynamics of the MLL1 complex on the NCP. We show that DPY30 binding to ASH2L leads to stabilization and integration of ASH2L IDRs into the MLL1 complex and establishes new ASH2L-NCP contacts. The significance of ASH2L-DPY30 interactions is demonstrated by requirement of both ASH2L IDRs and DPY30 for dramatic increase of processivity and activity of the MLL1 complex. This DPY30 and ASH2L-IDR dependent regulation is NCP-specific and applies to all members of the MLL/SET1 family of enzymes. We further show that DPY30 is causal for de novo establishment of H3K4me3 in ESCs. Our study provides a paradigm of how H3K4me3 is regulated on chromatin and how H3K4me3 heterogeneity can be modulated by ASH2L IDR interacting proteins.
- Department of Pathology, University of Michigan, Ann Arbor, MI, USA.
Organizational Affiliation: