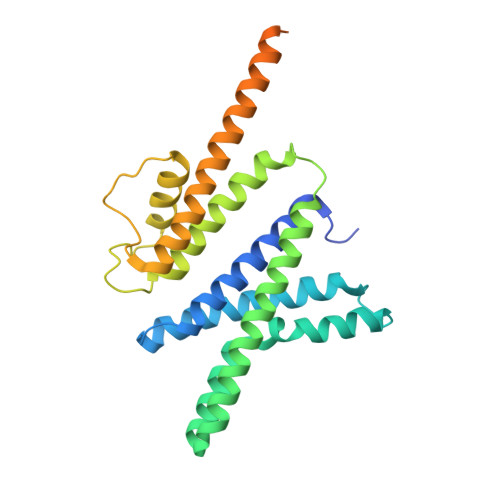
Cryo-EM structure of the KvAP channel reveals a non-domain-swapped voltage sensor topology.
Tao, X., MacKinnon, R.(2019) Elife 8
- PubMed: 31755864 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.52164
- Primary Citation Related Structures:
6UWM - PubMed Abstract:
Conductance in voltage-gated ion channels is regulated by membrane voltage through structural domains known as voltage sensors. A single structural class of voltage sensor domain exists, but two different modes of voltage sensor attachment to the pore occur in nature: domain-swapped and non-domain-swapped. Since the more thoroughly studied Kv1-7, Nav and Cav channels have domain-swapped voltage sensors, much less is known about non-domain-swapped voltage-gated ion channels. In this paper, using cryo-EM, we show that KvAP from Aeropyrum pernix has non-domain-swapped voltage sensors as well as other unusual features. The new structure, together with previous functional data, suggests that KvAP and the Shaker channel, to which KvAP is most often compared, probably undergo rather different voltage-dependent conformational changes when they open.
- Laboratory of Molecular Neurobiology and Biophysics, The Rockefeller University, Howard Hughes Medical Institute, New York, United States.
Organizational Affiliation: