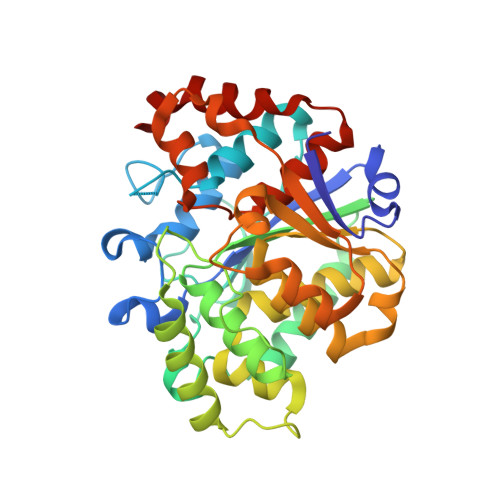
Structural studies of the phosphoribosyltransferase involved in cobamide biosynthesis in methanogenic archaea and cyanobacteria.
Jeter, V.L., Schwarzwalder, A.H., Rayment, I., Escalante-Semerena, J.C.(2022) Sci Rep 12: 17175-17175
- PubMed: 36229494 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-022-21765-5
- Primary Citation Related Structures:
6PT8, 6PTF, 6PU6 - PubMed Abstract:
Cobamides (Cbas) are coenzymes used by cells across all domains of life, but de novo synthesis is only found in some bacteria and archaea. Five enzymes assemble the nucleotide loop in the alpha phase of the corrin ring. Condensation of the activated ring and nucleobase yields adenosyl-Cba 5'-phosphate, which upon dephosphorylation yields the biologically active coenzyme (AdoCba). Base activation is catalyzed by a phosphoribosyltransferase (PRTase). The structure of the Salmonella enterica PRTase enzyme (i.e., SeCobT) is well-characterized, but archaeal PRTases are not. To gain insights into the mechanism of base activation by the PRTase from Methanocaldococcus jannaschii (MjCobT), we solved crystal structures of the enzyme in complex with substrate and products. We determined several structures: (i) a 2.2 Å structure of MjCobT in the absence of ligand (apo), (ii) structures of MjCobT bound to nicotinate mononucleotide (NaMN) and α-ribazole 5'-phosphate (α-RP) or α-adenylyl-5'-phosphate (α-AMP) at 2.3 and 1.4 Å, respectively. In MjCobT the general base that triggers the reaction is an aspartate residue (Asp 52) rather than a glutamate residue (E317) as in SeCobT. Notably, the dimer interface in MjCobT is completely different from that observed in SeCobT. Finally, entry PDB 3L0Z does not reflect the correct structure of MjCobT.
- Department of Microbiology, University of Georgia, Athens, GA, 30602, USA.
Organizational Affiliation: