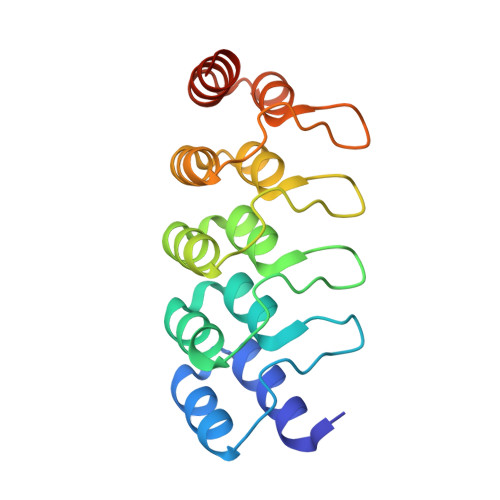
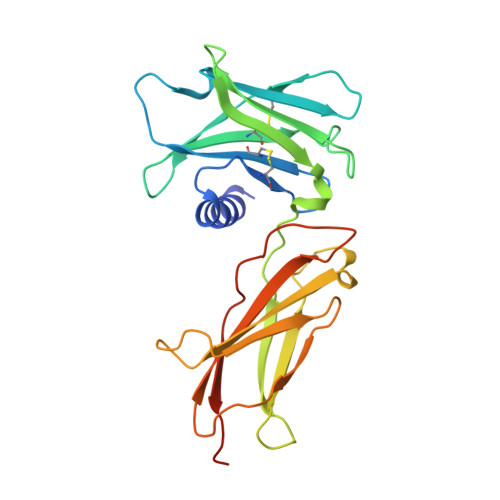
Topological control of cytokine receptor signaling induces differential effects in hematopoiesis.
Mohan, K., Ueda, G., Kim, A.R., Jude, K.M., Fallas, J.A., Guo, Y., Hafer, M., Miao, Y., Saxton, R.A., Piehler, J., Sankaran, V.G., Baker, D., Garcia, K.C.(2019) Science 364
- PubMed: 31123111 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aav7532
- Primary Citation Related Structures:
6MOE, 6MOF, 6MOG, 6MOH, 6MOI, 6MOJ, 6MOK, 6MOL - PubMed Abstract:
Although tunable signaling by G protein-coupled receptors can be exploited through medicinal chemistry, a comparable pharmacological approach has been lacking for the modulation of signaling through dimeric receptors, such as those for cytokines. We present a strategy to modulate cytokine receptor signaling output by use of a series of designed C2-symmetric cytokine mimetics, based on the designed ankyrin repeat protein (DARPin) scaffold, that can systematically control erythropoietin receptor (EpoR) dimerization orientation and distance between monomers. We sampled a range of EpoR geometries by varying intermonomer angle and distance, corroborated by several ligand-EpoR complex crystal structures. Across the range, we observed full, partial, and biased agonism as well as stage-selective effects on hematopoiesis. This surrogate ligand strategy opens access to pharmacological modulation of therapeutically important cytokine and growth factor receptor systems.
- Department of Molecular and Cellular Physiology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: