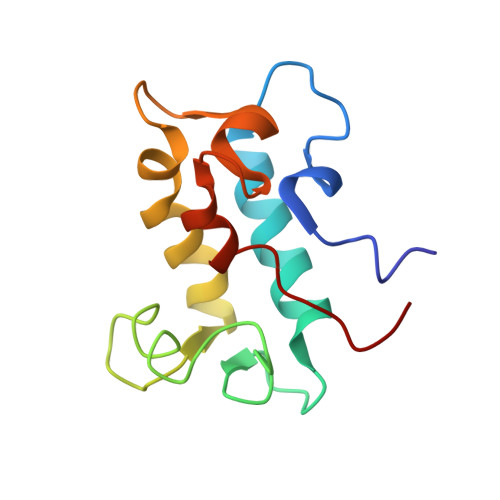
Structural Basis for Rare Earth Element Recognition by Methylobacterium extorquens Lanmodulin.
Cook, E.C., Featherston, E.R., Showalter, S.A., Cotruvo Jr., J.A.(2019) Biochemistry 58: 120-125
- PubMed: 30352145 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.8b01019
- Primary Citation Related Structures:
6MI5 - PubMed Abstract:
Lanmodulin (LanM) is a high-affinity lanthanide (Ln)-binding protein recently identified in Methylobacterium extorquens, a bacterium that requires Lns for the function of at least two enzymes. LanM possesses four EF-hands, metal coordination motifs generally associated with Ca II binding, but it undergoes a metal-dependent conformational change with a 100 million-fold selectivity for Ln III s and Y III over Ca II . Here we present the nuclear magnetic resonance solution structure of LanM complexed with Y III . This structure reveals that LanM features an unusual fusion of adjacent EF-hands, resulting in a compact fold to the best of our knowledge unique among EF-hand-containing proteins. It also supports the importance of an additional carboxylate ligand in contributing to the protein's picomolar affinity for Ln III s, and it suggests a role of unusual N i+1 -H···N i hydrogen bonds, in which LanM's unique EF-hand proline residues are engaged, in selective Ln III recognition. This work sets the stage for a detailed mechanistic understanding of LanM's Ln selectivity, which may inspire new strategies for binding, detecting, and sequestering these technologically important metals.
- Department of Chemistry , The Pennsylvania State University , University Park , Pennsylvania 16802 , United States.
Organizational Affiliation: