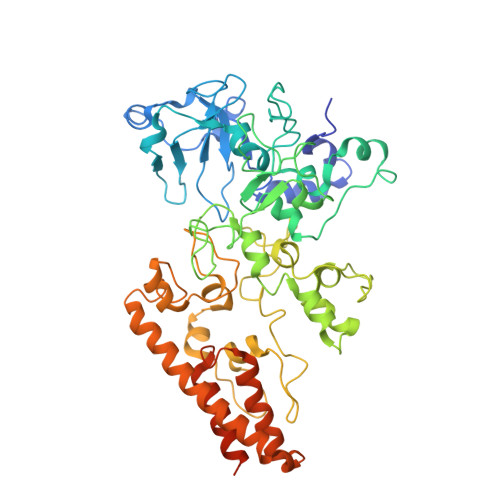
How Thermophilic Gram-Positive Organisms Perform Extracellular Electron Transfer: Characterization of the Cell Surface Terminal Reductase OcwA.
Costa, N.L., Hermann, B., Fourmond, V., Faustino, M.M., Teixeira, M., Einsle, O., Paquete, C.M., Louro, R.O.(2019) mBio 10
- PubMed: 31431546 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/mBio.01210-19
- Primary Citation Related Structures:
6I5B - PubMed Abstract:
Extracellular electron transfer is the key process underpinning the development of bioelectrochemical systems for the production of energy or added-value compounds. Thermincola potens JR is a promising Gram-positive bacterium to be used in these systems because it is thermophilic. In this paper, we describe the structural and functional properties of the nonaheme cytochrome OcwA, which is the terminal reductase of this organism. The structure of OcwA, determined at 2.2-Å resolution, shows that the overall fold and organization of the hemes are not related to other metal reductases and instead are similar to those of multiheme cytochromes involved in the biogeochemical cycles of nitrogen and sulfur. We show that, in addition to solid electron acceptors, OcwA can also reduce soluble electron shuttles and oxyanions. These data reveal that OcwA can work as a multipurpose respiratory enzyme allowing this organism to grow in environments with rapidly changing availability of terminal electron acceptors without the need for transcriptional regulation and protein synthesis. IMPORTANCE Thermophilic Gram-positive organisms were recently shown to be a promising class of organisms to be used in bioelectrochemical systems for the production of electrical energy. These organisms present a thick peptidoglycan layer that was thought to preclude them to perform extracellular electron transfer (i.e., exchange catabolic electrons with solid electron acceptors outside the cell). In this paper, we describe the structure and functional mechanisms of the multiheme cytochrome OcwA, the terminal reductase of the Gram-positive bacterium Thermincola potens JR found at the cell surface of this organism. The results presented here show that this protein can take the role of a respiratory "Swiss Army knife," allowing this organism to grow in environments with soluble and insoluble substrates. Moreover, it is shown that it is unrelated to terminal reductases found at the cell surface of other electroactive organisms. Instead, OcwA is similar to terminal reductases of soluble electron acceptors. Our data reveal that terminal oxidoreductases of soluble and insoluble substrates are evolutionarily related, providing novel insights into the evolutionary pathway of multiheme cytochromes.
- Instituto de Tecnologia Química e Biológica António Xavier, Universidade NOVA de Lisboa, Lisbon, Portugal.
Organizational Affiliation: