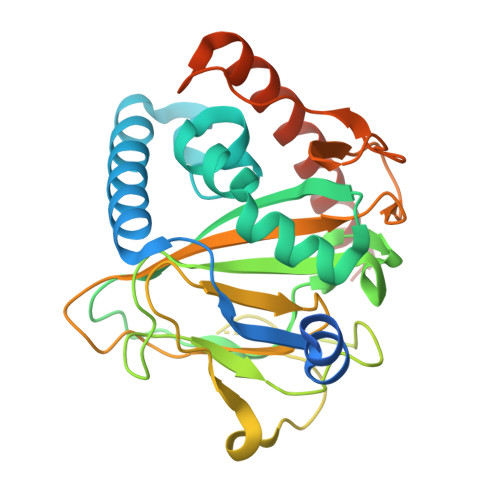
Non-Heme Monooxygenase ThoJ Catalyzes Thioholgamide beta-Hydroxylation.
Sikandar, A., Lopatniuk, M., Luzhetskyy, A., Koehnke, J.(2020) ACS Chem Biol 15: 2815-2819
- PubMed: 32965102 Search on PubMed
- DOI: https://doi.org/10.1021/acschembio.0c00637
- Primary Citation Related Structures:
6ZYK, 6ZYL - PubMed Abstract:
Thioviridamide-like compounds, including thioholgamides, are ribosomally synthesized and post-translationally modified peptide natural products with potent anticancer cell activity and an unprecedented structure. Very little is known about their biosynthesis, and we were intrigued by the β-hydroxy-N1, N3-dimethylhistidinium moiety found in these compounds. Here we report the construction of a heterologous host capable of producing thioholgamide with a 15-fold increased yield compared to the wild-type strain. A knockout of thoJ , encoding a predicted nonheme monooxygenase, shows that ThoJ is essential for thioholgamide β-hydroxylation. The crystal structure of ThoJ exhibits a typical mono/dioxygenase fold with conserved key active-site residues. Yet, ThoJ possesses a very large substrate binding pocket that appears suitable to receive a cyclic thioholgamide intermediate for hydroxylation. The improved production of the heterologous host will enable the dissection of the individual biosynthetic steps involved in biosynthesis of this exciting RiPP family.
- Workgroup Structural Biology of Biosynthetic Enzymes, Helmholtz Institute for Pharmaceutical Research Saarland (HIPS), Helmholtz Centre for Infection Research (HZI), Saarland University, Campus Geb. E8.1, 66123 Saarbrücken, Germany.
Organizational Affiliation: