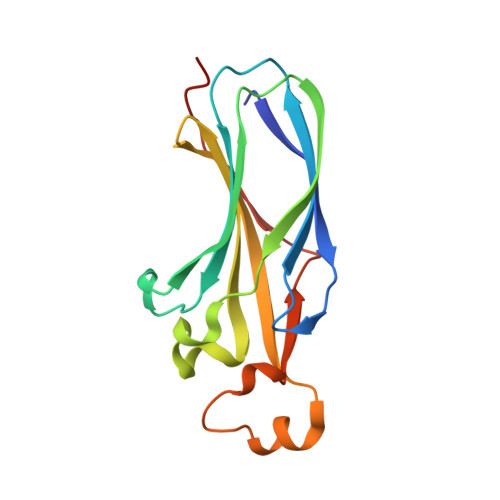
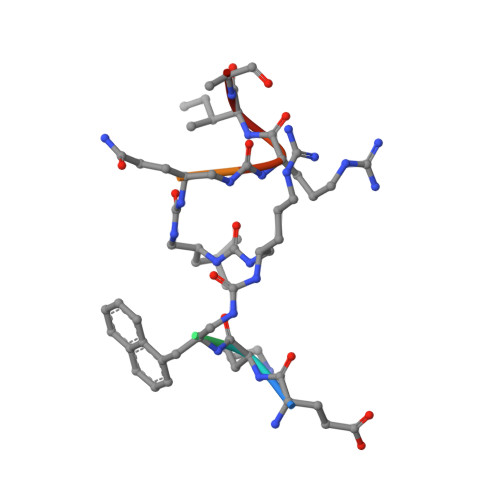
Optimal anchoring of a foldamer inhibitor of ASF1 histone chaperone through backbone plasticity.
Mbianda, J., Bakail, M., Andre, C., Moal, G., Perrin, M.E., Pinna, G., Guerois, R., Becher, F., Legrand, P., Traore, S., Douat, C., Guichard, G., Ochsenbein, F.(2021) Sci Adv 7
- PubMed: 33741589 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abd9153
- Primary Citation Related Structures:
6ZUF - PubMed Abstract:
Sequence-specific oligomers with predictable folding patterns, i.e., foldamers, provide new opportunities to mimic α-helical peptides and design inhibitors of protein-protein interactions. One major hurdle of this strategy is to retain the correct orientation of key side chains involved in protein surface recognition. Here, we show that the structural plasticity of a foldamer backbone may notably contribute to the required spatial adjustment for optimal interaction with the protein surface. By using oligoureas as α helix mimics, we designed a foldamer/peptide hybrid inhibitor of histone chaperone ASF1, a key regulator of chromatin dynamics. The crystal structure of its complex with ASF1 reveals a notable plasticity of the urea backbone, which adapts to the ASF1 surface to maintain the same binding interface. One additional benefit of generating ASF1 ligands with nonpeptide oligourea segments is the resistance to proteolysis in human plasma, which was highly improved compared to the cognate α-helical peptide.
- Univ. Bordeaux, CNRS, Bordeaux INP, CBMN, UMR 5248, Institut Européen de Chimie et Biologie, 2 rue Robert Escarpit, F-33607 Pessac, France.
Organizational Affiliation: