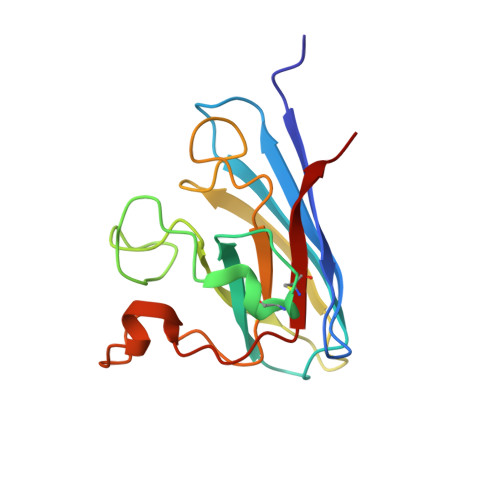
Crystal Structure of a Cu,Zn Superoxide Dismutase From the Thermophilic Fungus Chaetomium thermophilum.
Mohsin, I., Zhang, L.Q., Li, D.C., Papageorgiou, A.C.(2021) Protein Pept Lett 28: 1043-1053
- PubMed: 33726638 Search on PubMed
- DOI: https://doi.org/10.2174/0929866528666210316104919
- Primary Citation Related Structures:
6ZS1 - PubMed Abstract:
Thermophilic fungi have recently emerged as a promising source of thermostable enzymes. Superoxide dismutases are key antioxidant metalloenzymes with promising therapeutic effects in various diseases, both acute and chronic. However, structural heterogeneity and low thermostability limit their therapeutic efficacy. Although several studies from hypethermophilic superoxide dismutases (SODs) have been reported, information about Cu,Zn-SODs from thermophilic fungi is scarce. Chaetomium thermophilum is a thermophilic fungus that could provide proteins with thermophilic properties. The enzyme was expressed in Pichia pastoris cells and crystallized using the vapor-diffusion method. X-ray data were collected, and the structure was determined and refined to 1.56 Å resolution. Structural analysis and comparisons were carried out. The presence of 8 molecules (A through H) in the asymmetric unit resulted in four different interfaces. Molecules A and F form the typical homodimer which is also found in other Cu,Zn- SODs. Zinc was present in all subunits of the structure while copper was found in only four subunits with reduced occupancy (C, D, E and F). The ability of the enzyme to form oligomers and the elevated Thr:Ser ratio may be contributing factors to its thermal stability. Two hydrophobic residues that participate in interface formation and are not present in other CuZn-SODs may play a role in the formation of new interfaces and the oligomerization process. The CtSOD crystal structure reported here is the first Cu,Zn-SOD structure from a thermophilic fungus.
- Turku Bioscience Centre, University of Turku and Åbo Akademi University, Turku 20521, Finland.
Organizational Affiliation: