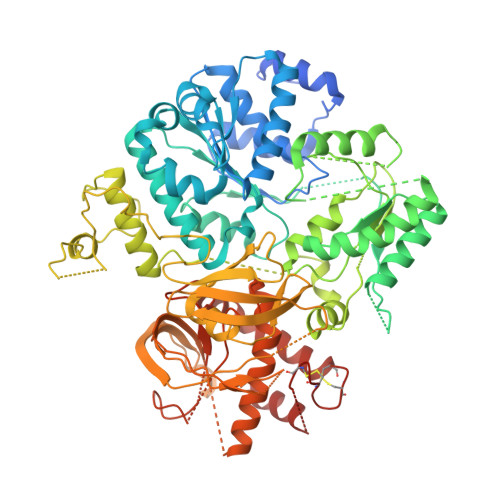
A skipping rope translocation mechanism in a widespread family of DNA repair helicases.
Roske, J.J., Liu, S., Loll, B., Neu, U., Wahl, M.C.(2021) Nucleic Acids Res 49: 504-518
- PubMed: 33300032 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa1174
- Primary Citation Related Structures:
6ZNP, 6ZNQ, 6ZNS - PubMed Abstract:
Mitomycin repair factor A represents a family of DNA helicases that harbor a domain of unknown function (DUF1998) and support repair of mitomycin C-induced DNA damage by presently unknown molecular mechanisms. We determined crystal structures of Bacillus subtilis Mitomycin repair factor A alone and in complex with an ATP analog and/or DNA and conducted structure-informed functional analyses. Our results reveal a unique set of auxiliary domains appended to a dual-RecA domain core. Upon DNA binding, a Zn2+-binding domain, encompassing the domain of unknown function, acts like a drum that rolls out a canopy of helicase-associated domains, entrapping the substrate and tautening an inter-domain linker across the loading strand. Quantification of DNA binding, stimulated ATPase and helicase activities in the wild type and mutant enzyme variants in conjunction with the mode of coordination of the ATP analog suggest that Mitomycin repair factor A employs similar ATPase-driven conformational changes to translocate on DNA, with the linker ratcheting through the nucleotides like a 'skipping rope'. The electrostatic surface topology outlines a likely path for the displaced DNA strand. Our results reveal unique molecular mechanisms in a widespread family of DNA repair helicases linked to bacterial antibiotics resistance.
- Freie Universität Berlin, Institute of Chemistry and Biochemistry, Laboratory of Structural Biochemistry, Takustraβe 6, D-14195 Berlin, Germany.
Organizational Affiliation: