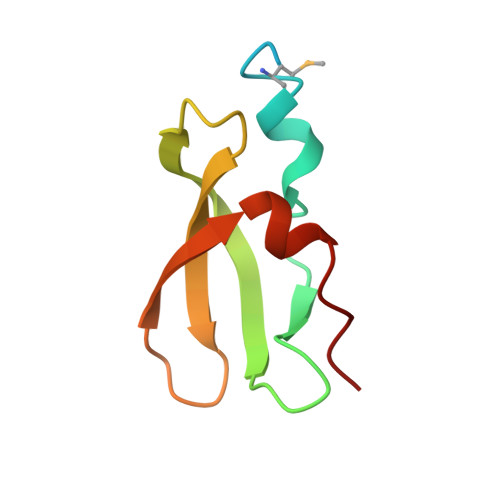
Structural Basis for Toxin Inhibition in the VapXD Toxin-Antitoxin System.
Bertelsen, M.B., Senissar, M., Nielsen, M.H., Bisiak, F., Cunha, M.V., Molinaro, A.L., Daines, D.A., Brodersen, D.E.(2021) Structure 29: 139
- PubMed: 33096014 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2020.10.002
- Primary Citation Related Structures:
6ZI0, 6ZI1, 6ZN8 - PubMed Abstract:
Bacterial type II toxin-antitoxin (TA) modules encode a toxic protein that downregulates metabolism and a specific antitoxin that binds and inhibits the toxin during normal growth. In non-typeable Haemophilus influenzae, a common cause of infections in humans, the vapXD locus was found to constitute a functional TA module and contribute to pathogenicity; however, the mode of action of VapD and the mechanism of inhibition by the VapX antitoxin remain unknown. Here, we report the structure of the intact H. influenzae VapXD complex, revealing an unusual 2:1 TA molecular stoichiometry where a Cas2-like homodimer of VapD binds a single VapX antitoxin. VapX consists of an oligonucleotide/oligosaccharide-binding domain that docks into an asymmetrical cavity on the toxin dimer. Structures of isolated VapD further reveal how a symmetrical toxin homodimer adapts to interacting with an asymmetrical antitoxin and suggest how a primordial TA system evolved to become part of CRISPR-Cas immunity systems.
- Department of Molecular Biology and Genetics, Aarhus University, DK-8000 Aarhus C, Denmark.
Organizational Affiliation: