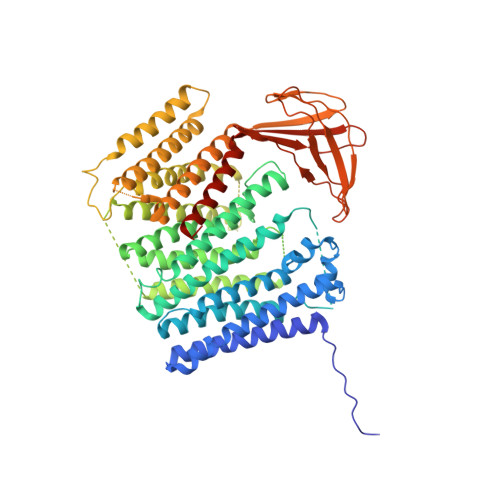
Architecture of the membrane-bound cytochrome c heme lyase CcmF.
Brausemann, A., Zhang, L., Ilcu, L., Einsle, O.(2021) Nat Chem Biol 17: 800-805
- PubMed: 33958791 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-021-00793-8
- Primary Citation Related Structures:
6ZMQ - PubMed Abstract:
The covalent attachment of one or multiple heme cofactors to cytochrome c protein chains enables cytochrome c proteins to be used in electron transfer and redox catalysis in extracytoplasmic environments. A dedicated heme maturation machinery, whose core component is a heme lyase, scans nascent peptides after Sec-dependent translocation for CX n CH-binding motifs. Here we report the three-dimensional (3D) structure of the heme lyase CcmF, a 643-amino acid integral membrane protein, from Thermus thermophilus. CcmF contains a heme b cofactor at the bottom of a large cavity that opens toward the extracellular side to receive heme groups from the heme chaperone CcmE for cytochrome maturation. A surface groove on CcmF may guide the extended apoprotein to heme attachment at or near a loop containing the functionally essential WXWD motif, which is situated above the putative cofactor binding pocket. The structure suggests heme delivery from within the membrane, redefining the role of the chaperone CcmE.
- Institut für Biochemie, Albert-Ludwigs-Universität Freiburg, Freiburg, Germany.
Organizational Affiliation: