In vitro reconstitution of dynamically interacting integral membrane subunits of energy-coupling factor transporters.
Setyawati, I., Stanek, W.K., Majsnerowska, M., Swier, L.J.Y.M., Pardon, E., Steyaert, J., Guskov, A., Slotboom, D.J.(2020) Elife 9
- PubMed: 33350937 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.64389
- Primary Citation Related Structures:
6ZG3 - PubMed Abstract:
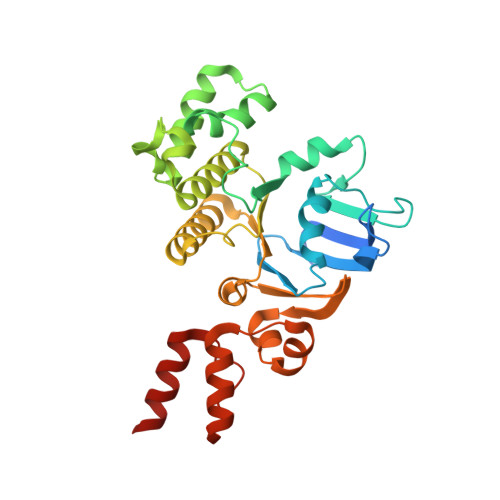
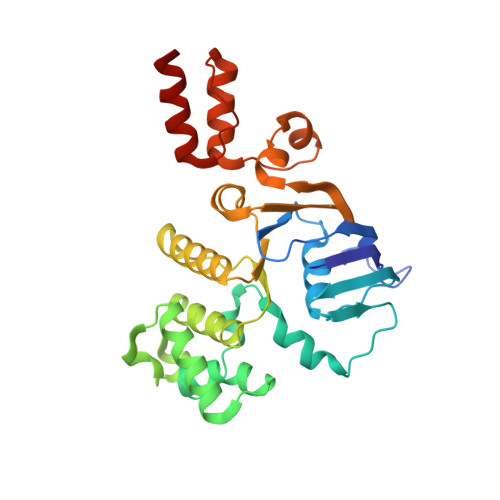
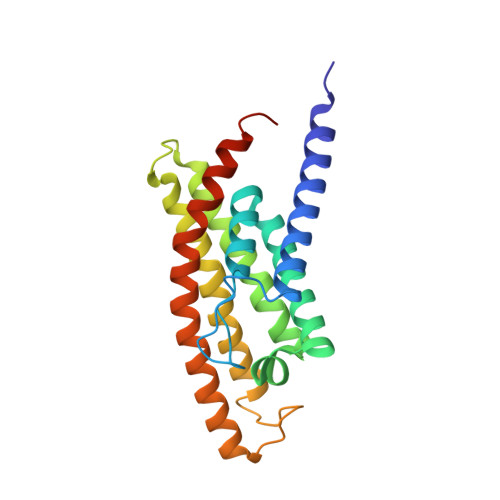
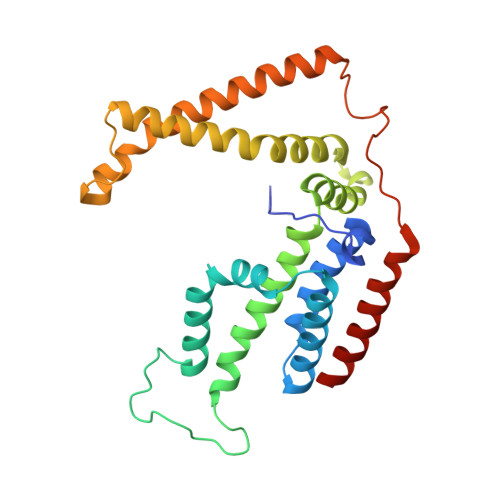
Energy-coupling factor (ECF) transporters mediate import of micronutrients in prokaryotes. They consist of an integral membrane S-component (that binds substrate) and ECF module (that powers transport by ATP hydrolysis). It has been proposed that different S-components compete for docking onto the same ECF module, but a minimal liposome-reconstituted system, required to substantiate this idea, is lacking. Here, we co-reconstituted ECF transporters for folate (ECF-FolT2) and pantothenate (ECF-PanT) into proteoliposomes, and assayed for crosstalk during active transport. The kinetics of transport showed that exchange of S-components is part of the transport mechanism. Competition experiments suggest much slower substrate association with FolT2 than with PanT. Comparison of a crystal structure of ECF-PanT with previously determined structures of ECF-FolT2 revealed larger conformational changes upon binding of folate than pantothenate, which could explain the kinetic differences. Our work shows that a minimal in vitro system with two reconstituted transporters recapitulates intricate kinetics behaviour observed in vivo.
- Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Groningen, Netherlands.
Organizational Affiliation: