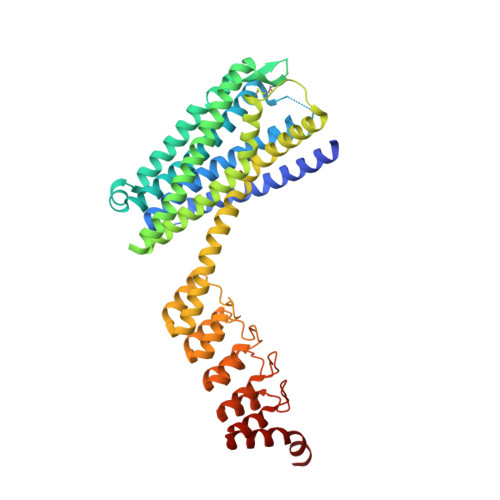
Complexes of the neurotensin receptor 1 with small-molecule ligands reveal structural determinants of full, partial, and inverse agonism.
Deluigi, M., Klipp, A., Klenk, C., Merklinger, L., Eberle, S.A., Morstein, L., Heine, P., Mittl, P.R.E., Ernst, P., Kamenecka, T.M., He, Y., Vacca, S., Egloff, P., Honegger, A., Pluckthun, A.(2021) Sci Adv 7
- PubMed: 33571132 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abe5504
- Primary Citation Related Structures:
6YVR, 6Z4Q, 6Z4S, 6Z4V, 6Z66, 6Z8N, 6ZA8, 6ZIN - PubMed Abstract:
Neurotensin receptor 1 (NTSR1) and related G protein-coupled receptors of the ghrelin family are clinically unexploited, and several mechanistic aspects of their activation and inactivation have remained unclear. Enabled by a new crystallization design, we present five new structures: apo-state NTSR1 as well as complexes with nonpeptide inverse agonists SR48692 and SR142948A, partial agonist RTI-3a, and the novel full agonist SRI-9829, providing structural rationales on how ligands modulate NTSR1. The inverse agonists favor a large extracellular opening of helices VI and VII, undescribed so far for NTSR1, causing a constriction of the intracellular portion. In contrast, the full and partial agonists induce a binding site contraction, and their efficacy correlates with the ability to mimic the binding mode of the endogenous agonist neurotensin. Providing evidence of helical and side-chain rearrangements modulating receptor activation, our structural and functional data expand the mechanistic understanding of NTSR1 and potentially other peptidergic receptors.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland.
Organizational Affiliation: