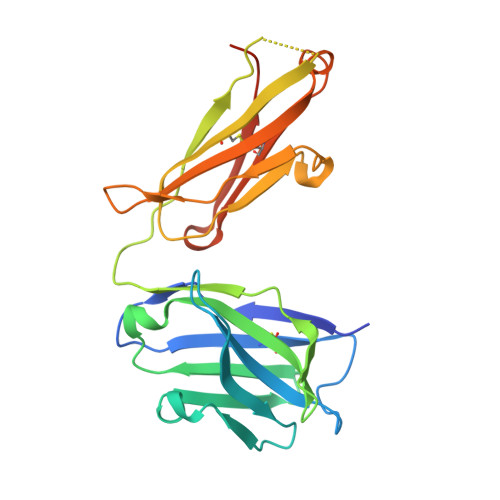
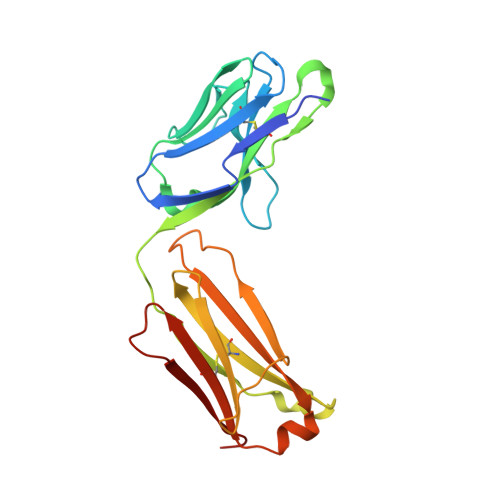
Structural and functional analysis of protective antibodies targeting the threefold plateau of enterovirus 71.
Huang, K.A., Zhou, D., Fry, E.E., Kotecha, A., Huang, P.N., Yang, S.L., Tsao, K.C., Huang, Y.C., Lin, T.Y., Ren, J., Stuart, D.I.(2020) Nat Commun 11: 5253-5253
- PubMed: 33067459 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19013-3
- Primary Citation Related Structures:
6Z3K, 6Z3P, 6Z3Q - PubMed Abstract:
Enterovirus 71 (EV71)-neutralizing antibodies correlate with protection and have potential as therapeutic agents. We isolate and characterize a panel of plasmablast-derived monoclonal antibodies from an infected child whose antibody response focuses on the plateau epitope near the icosahedral 3-fold axes. Eight of a total of 19 antibodies target this epitope and three of these potently neutralize the virus. Representative neutralizing antibodies 38-1-10A and 38-3-11A both confer effective protection against lethal EV71 challenge in hSCARB2-transgenic mice. The cryo-electron microscopy structures of the EV71 virion in complex with Fab fragments of these potent and protective antibodies reveal the details of a conserved epitope formed by residues in the BC and HI loops of VP2 and the BC and HI loops of VP3 spanning the region around the 3-fold axis. Remarkably, the two antibodies interact with the epitope in quite distinct ways. These plateau-binding antibodies provide templates for promising candidate therapeutics.
- Division of Pediatric Infectious Diseases, Department of Pediatrics, Chang Gung Memorial Hospital, Taoyuan, Taiwan. arthur1726@cgmh.org.tw.
Organizational Affiliation: