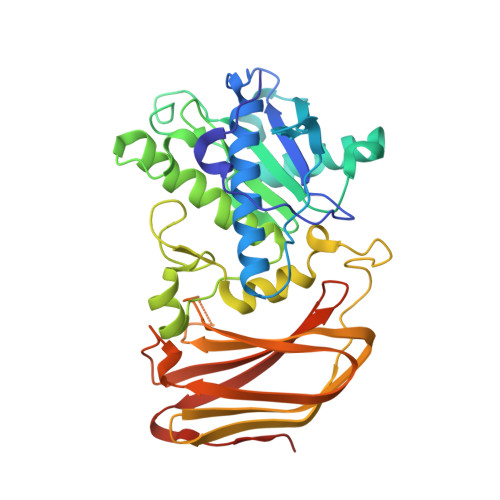
Structural basis of mammalian mucin processing by the human gut O-glycopeptidase OgpA from Akkermansia muciniphila.
Trastoy, B., Naegeli, A., Anso, I., Sjogren, J., Guerin, M.E.(2020) Nat Commun 11: 4844-4844
- PubMed: 32973204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-18696-y
- Primary Citation Related Structures:
6Z2D, 6Z2O, 6Z2P, 6Z2Q - PubMed Abstract:
Akkermansia muciniphila is a mucin-degrading bacterium commonly found in the human gut that promotes a beneficial effect on health, likely based on the regulation of mucus thickness and gut barrier integrity, but also on the modulation of the immune system. In this work, we focus in OgpA from A. muciniphila, an O-glycopeptidase that exclusively hydrolyzes the peptide bond N-terminal to serine or threonine residues substituted with an O-glycan. We determine the high-resolution X-ray crystal structures of the unliganded form of OgpA, the complex with the glycodrosocin O-glycopeptide substrate and its product, providing a comprehensive set of snapshots of the enzyme along the catalytic cycle. In combination with O-glycopeptide chemistry, enzyme kinetics, and computational methods we unveil the molecular mechanism of O-glycan recognition and specificity for OgpA. The data also contribute to understanding how A. muciniphila processes mucins in the gut, as well as analysis of post-translational O-glycosylation events in proteins.
- Structural Biology Unit, Center for Cooperative Research in Biosciences (CIC bioGUNE), Basque Research and Technology Alliance (BRTA), Bizkaia Technology Park, Building 801A, 48160, Derio, Spain.
Organizational Affiliation: