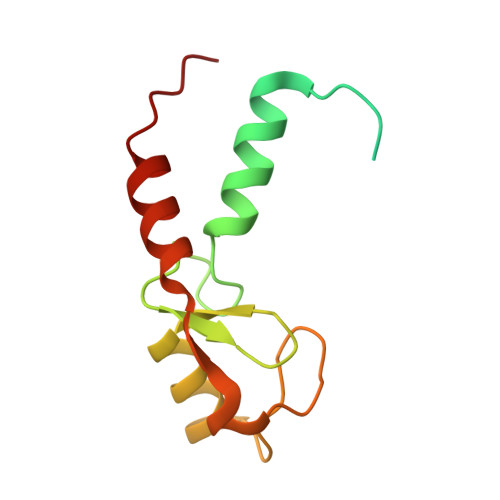
The RING domain of TRIM69 promotes higher-order assembly.
Keown, J.R., Yang, J., Black, M.M., Goldstone, D.C.(2020) Acta Crystallogr D Struct Biol 76: 954-961
- PubMed: 33021497 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798320010499
- Primary Citation Related Structures:
6YXE - PubMed Abstract:
Members of the TRIM protein family have been shown to inhibit a range of viral infections. Recently, TRIM69 was identified as a potent inhibitor of Vesicular stomatitis Indiana virus infection, with its inhibition being dependent upon multimerization. Using SEC-MALLS analysis, it is demonstrated that the assembly of TRIM69 is mediated through the RING domain and not the Bbox domain as has been shown for other TRIM proteins. Using X-ray crystallography, the structure of the TRIM69 RING domain has been determined to a resolution of 2.1 Å, the oligomerization interface has been identified and regions outside the four-helix bundle have been observed to form interactions that are likely to support assembly.
- School of Biological Sciences, University of Auckland, Private Bag 92019, Auckland, New Zealand.
Organizational Affiliation: