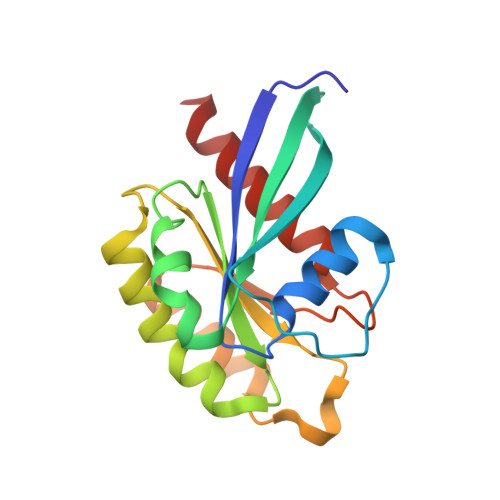
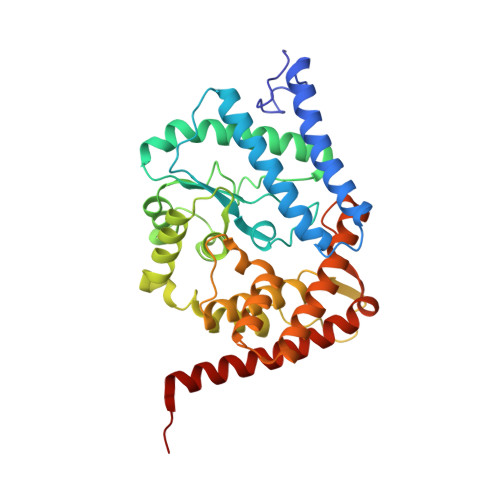
Rab1-AMPylation by Legionella DrrA is allosterically activated by Rab1.
Du, J., Wrisberg, M.V., Gulen, B., Stahl, M., Pett, C., Hedberg, C., Lang, K., Schneider, S., Itzen, A.(2021) Nat Commun 12: 460-460
- PubMed: 33469029 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-20702-2
- Primary Citation Related Structures:
6YX5 - PubMed Abstract:
Legionella pneumophila infects eukaryotic cells by forming a replicative organelle - the Legionella containing vacuole. During this process, the bacterial protein DrrA/SidM is secreted and manipulates the activity and post-translational modification (PTM) states of the vesicular trafficking regulator Rab1. As a result, Rab1 is modified with an adenosine monophosphate (AMP), and this process is referred to as AMPylation. Here, we use a chemical approach to stabilise low-affinity Rab:DrrA complexes in a site-specific manner to gain insight into the molecular basis of the interaction between the Rab protein and the AMPylation domain of DrrA. The crystal structure of the Rab:DrrA complex reveals a previously unknown non-conventional Rab-binding site (NC-RBS). Biochemical characterisation demonstrates allosteric stimulation of the AMPylation activity of DrrA via Rab binding to the NC-RBS. We speculate that allosteric control of DrrA could in principle prevent random and potentially cytotoxic AMPylation in the host, thereby perhaps ensuring efficient infection by Legionella.
- Center for Integrated Protein Science Munich (CIPSM), Department of Chemistry, Technical University of Munich, Garching, 85748, Germany.
Organizational Affiliation: