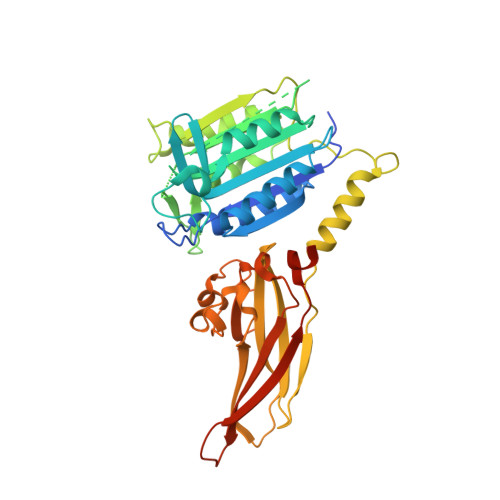
Stabilizing Inactive Conformations of MALT1 as an Effective Approach to Inhibit Its Protease Activity
Hughes, N., Erbel, P., Bornancin, F., Wiesmann, C., Schiering, N., Villard, F., Decock, A., Rubi, B., Melkko, S., Spanka, C., Buschmann, N., Pissot-Soldermann, C., Simic, O., Beerli, R., Sorge, M., Tintelnot-Blomley, M., Beltz, K., Regnier, C.H., Quancard, J., Schlapbach, A., Langlois, J., Renatus, M.(2020) Adv Ther (Weinh) 3