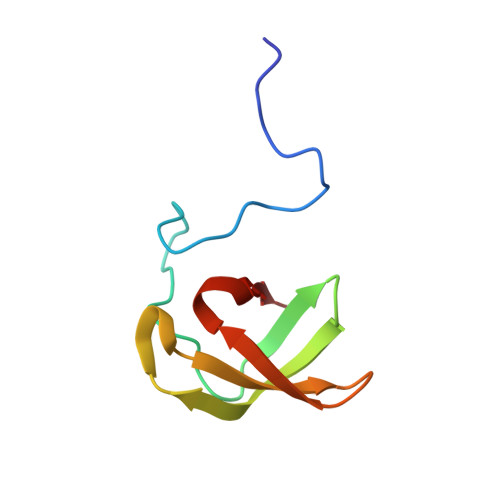
UvrD helicase-RNA polymerase interactions are governed by UvrD's carboxy-terminal Tudor domain.
Kawale, A.A., Burmann, B.M.(2020) Commun Biol 3: 607-607
- PubMed: 33097771 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-01332-2
- Primary Citation Related Structures:
6YHZ, 6YI2 - PubMed Abstract:
All living organisms have to cope with the constant threat of genome damage by UV light and other toxic reagents. To maintain the integrity of their genomes, organisms developed a variety of DNA repair pathways. One of these, the Transcription Coupled DNA-Repair (TCR) pathway, is triggered by stalled RNA Polymerase (RNAP) complexes at DNA damage sites on actively transcribed genes. A recently elucidated bacterial TCR pathway employs the UvrD helicase pulling back stalled RNAP complexes from the damage, stimulating recruitment of the DNA-repair machinery. However, structural and functional aspects of UvrD's interaction with RNA Polymerase remain elusive. Here we used advanced solution NMR spectroscopy to investigate UvrD's role within the TCR, identifying that the carboxy-terminal region of the UvrD helicase facilitates RNAP interactions by adopting a Tudor-domain like fold. Subsequently, we functionally analyzed this domain, identifying it as a crucial component for the UvrD-RNAP interaction besides having nucleic-acid affinity.
- Wallenberg Centre for Molecular and Translational Medicine, University of Gothenburg, Gothenburg, 40530, Sweden.
Organizational Affiliation: