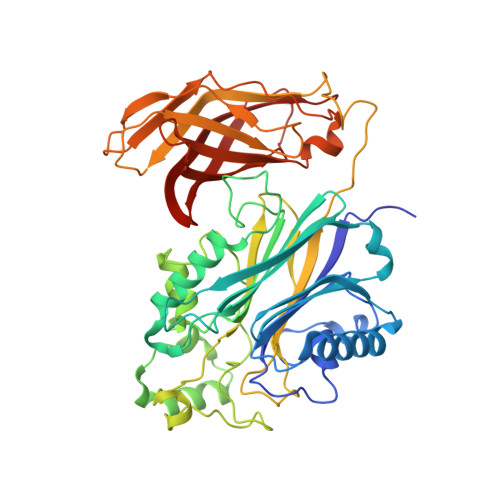
Regulation of inositol 5-phosphatase activity by the C2 domain of SHIP1 and SHIP2.
Bradshaw, W.J., Kennedy, E.C., Moreira, T., Smith, L.A., Chalk, R., Katis, V.L., Benesch, J.L.P., Brennan, P.E., Murphy, E.J., Gileadi, O.(2024) Structure
- PubMed: 38309262 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2024.01.005
- Primary Citation Related Structures:
5RW2, 5RW3, 5RW4, 5RW5, 5RW6, 5RW7, 5RW8, 5RW9, 5RWA, 5RWB, 5RWC, 5RWD, 5RWE, 5RWF, 5RWG, 5RWH, 5RWI, 5RWJ, 5RWK, 5RWL, 5RWM, 5RWN, 5RWO, 5RWP, 5RWQ, 5RWR, 5RWS, 5RWT, 5RWU, 5RWV, 5RWW, 5RWX, 5RWY, 5RWZ, 5RX0, 5RX1, 5RX2, 5RX3, 5RX4, 5RX5, 5RX6, 5RX7, 5RX8, 5RX9, 5RXA, 5RXB, 5RXC, 5RXD, 5RXE, 5RXF, ... Search all related entries - PubMed Abstract:
SHIP1, an inositol 5-phosphatase, plays a central role in cellular signaling. As such, it has been implicated in many conditions. Exploiting SHIP1 as a drug target will require structural knowledge and the design of selective small molecules. We have determined apo, and magnesium and phosphate-bound structures of the phosphatase and C2 domains of SHIP1. The C2 domains of SHIP1 and the related SHIP2 modulate the activity of the phosphatase domain. To understand the mechanism, we performed activity assays, hydrogen-deuterium exchange mass spectrometry, and molecular dynamics on SHIP1 and SHIP2. Our findings demonstrate that the influence of the C2 domain is more pronounced for SHIP2 than SHIP1. We determined 91 structures of SHIP1 with fragments bound, with some near the interface between the two domains. We performed a mass spectrometry screen and determined four structures with covalent fragments. These structures could act as starting points for the development of potent, selective probes.
- ARUK Oxford Drug Discovery Institute, Centre for Medicines Discovery, Nuffield Department of Medicine Research Building, Old Road Campus, University of Oxford, Oxford OX3 7FZ, UK. Electronic address: william.bradshaw@cmd.ox.ac.uk.
Organizational Affiliation: