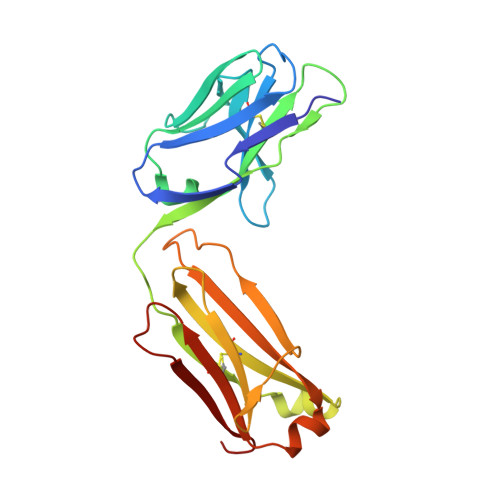
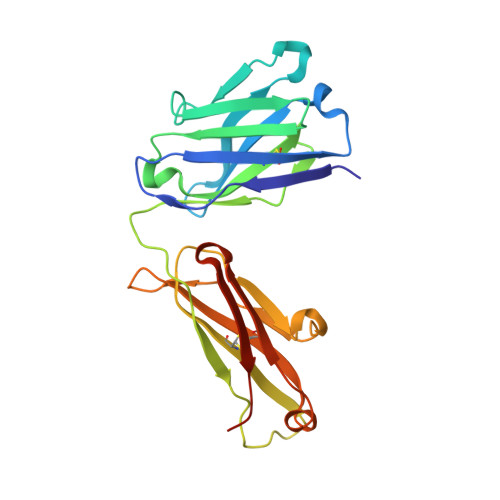
Biomolecular Recognition of the Glycan Neoantigen CA19-9 by Distinct Antibodies.
Borenstein-Katz, A., Warszawski, S., Amon, R., Eilon, M., Cohen-Dvashi, H., Leviatan Ben-Arye, S., Tasnima, N., Yu, H., Chen, X., Padler-Karavani, V., Fleishman, S.J., Diskin, R.(2021) J Mol Biology 433: 167099-167099
- PubMed: 34119488 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2021.167099
- Primary Citation Related Structures:
6XTG, 6XUD, 6XUK, 6XUL, 6XUN - PubMed Abstract:
Glycans decorate the cell surface, secreted glycoproteins and glycolipids, and altered glycans are often found in cancers. Despite their high diagnostic and therapeutic potential, however, glycans are polar and flexible molecules that are quite challenging for the development and design of high-affinity binding antibodies. To understand the mechanisms by which glycan neoantigens are specifically recognized by antibodies, we analyze the biomolecular recognition of the tumor-associated carbohydrate antigen CA19-9 by two distinct antibodies using X-ray crystallography. Despite the potential plasticity of glycans and the very different antigen-binding surfaces presented by the antibodies, both structures reveal an essentially identical extended CA19-9 conformer, suggesting that this conformer's stability selects the antibodies. Starting from the bound structure of one of the antibodies, we use the AbLIFT computational algorithm to design a variant with seven core mutations in the variable domain's light-heavy chain interface that exhibits tenfold improved affinity for CA19-9. The results reveal strategies used by antibodies to specifically recognize glycan antigens and show how automated antibody-optimization methods may be used to enhance the clinical potential of existing antibodies.
- Department of Chemical and Structural Biology, Weizmann Institute of Science, 76100 Rehovot, Israel.
Organizational Affiliation: