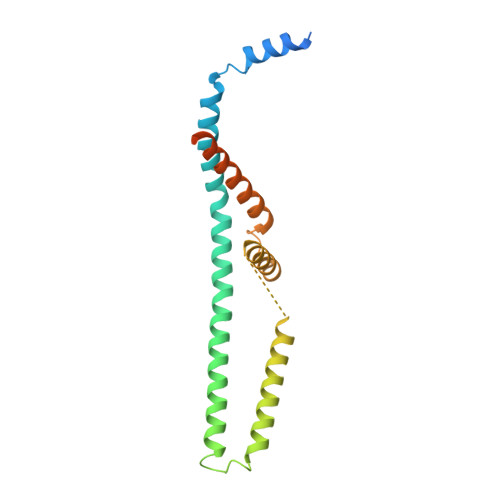
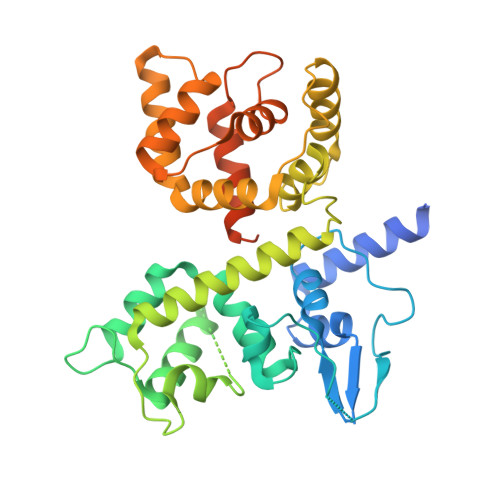
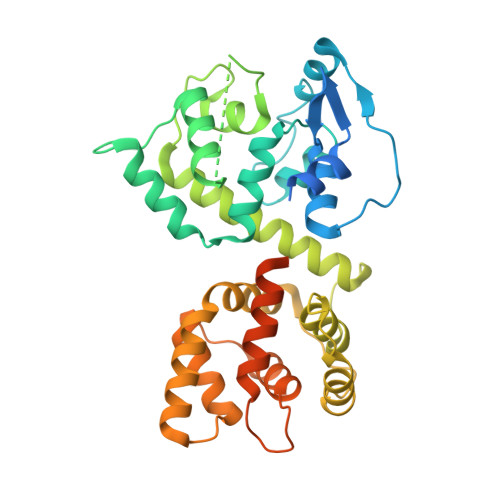
Structures reveal gatekeeping of the mitochondrial Ca 2+ uniporter by MICU1-MICU2.
Wang, C., Jacewicz, A., Delgado, B.D., Baradaran, R., Long, S.B.(2020) Elife 9
- PubMed: 32667285 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.59991
- Primary Citation Related Structures:
6XQN, 6XQO - PubMed Abstract:
The mitochondrial calcium uniporter is a Ca 2+ -gated ion channel complex that controls mitochondrial Ca 2+ entry and regulates cell metabolism. MCU and EMRE form the channel while Ca 2+ -dependent regulation is conferred by MICU1 and MICU2 through an enigmatic process. We present a cryo-EM structure of an MCU-EMRE-MICU1-MICU2 holocomplex comprising MCU and EMRE subunits from the beetle Tribolium castaneum in complex with a human MICU1-MICU2 heterodimer at 3.3 Å resolution. With analogy to how neuronal channels are blocked by protein toxins, a uniporter interaction domain on MICU1 binds to a channel receptor site comprising MCU and EMRE subunits to inhibit ion flow under resting Ca 2+ conditions. A Ca 2+ -bound structure of MICU1-MICU2 at 3.1 Å resolution indicates how Ca 2+ -dependent changes enable dynamic response to cytosolic Ca 2+ signals.
- Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, United States.
Organizational Affiliation: