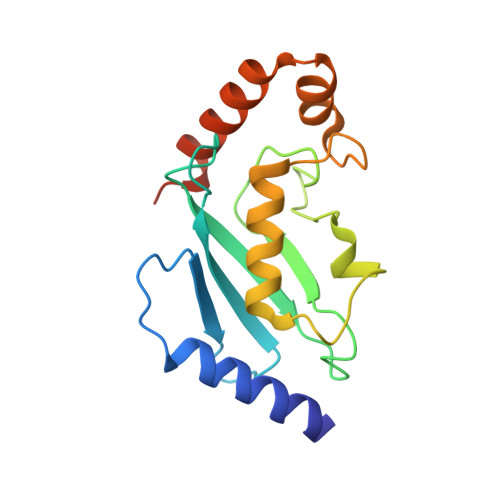
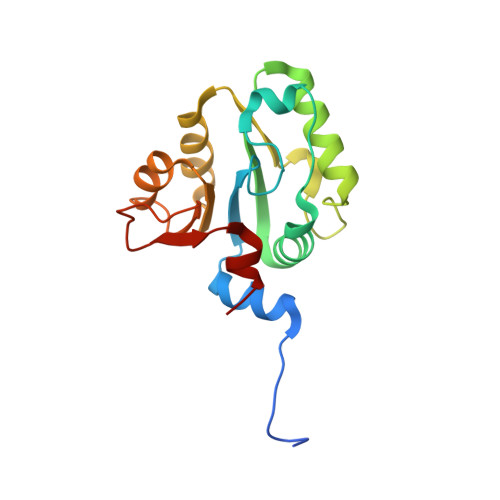
The Structure of the Arabidopsis PEX4-PEX22 Peroxin Complex-Insights Into Ubiquitination at the Peroxisomal Membrane
Traver, M.S., Bradford, S.E., Olmos, J.L., Wright, Z.J., Miller, M.D., Xu, W., Phillips, G.N., Bartel, B.(2022) Front Cell Dev Biol 10: 838923-838923