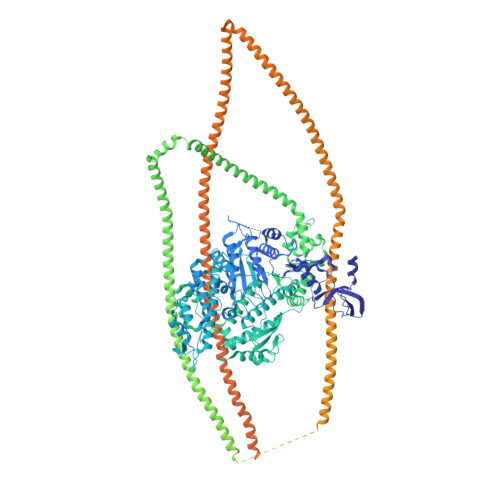
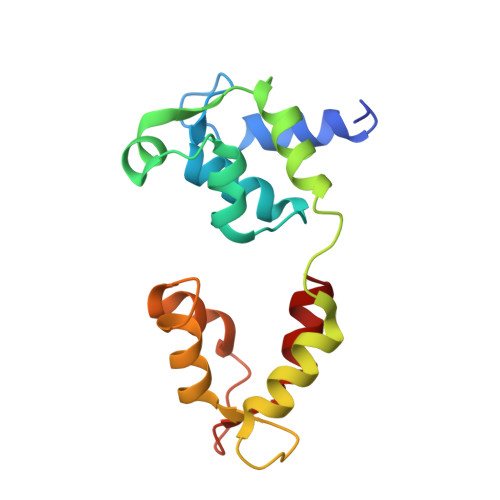
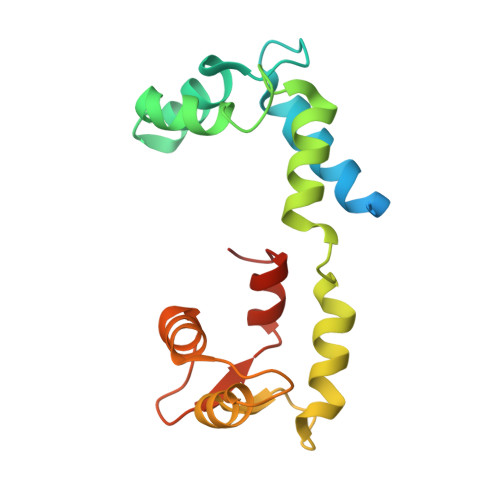
Cryo-EM structure of the inhibited (10S) form of myosin II.
Yang, S., Tiwari, P., Lee, K.H., Sato, O., Ikebe, M., Padron, R., Craig, R.(2020) Nature 588: 521-525
- PubMed: 33268893 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-3007-0
- Primary Citation Related Structures:
6XE9 - PubMed Abstract:
Myosin II is the motor protein that enables muscle cells to contract and nonmuscle cells to move and change shape 1 . The molecule has two identical heads attached to an elongated tail, and can exist in two conformations: 10S and 6S, named for their sedimentation coefficients 2,3 . The 6S conformation has an extended tail and assembles into polymeric filaments, which pull on actin filaments to generate force and motion. In 10S myosin, the tail is folded into three segments and the heads bend back and interact with each other and the tail 3-7 , creating a compact conformation in which ATPase activity, actin activation and filament assembly are all highly inhibited 7,8 . This switched-off structure appears to function as a key energy-conserving storage molecule in muscle and nonmuscle cells 9-12 , which can be activated to form functional filaments as needed 13 -but the mechanism of its inhibition is not understood. Here we have solved the structure of smooth muscle 10S myosin by cryo-electron microscopy with sufficient resolution to enable improved understanding of the function of the head and tail regions of the molecule and of the key intramolecular contacts that cause inhibition. Our results suggest an atomic model for the off state of myosin II, for its activation and unfolding by phosphorylation, and for understanding the clustering of disease-causing mutations near sites of intramolecular interaction.
- Division of Cell Biology and Imaging, Department of Radiology, University of Massachusetts Medical School, Worcester, MA, USA.
Organizational Affiliation: