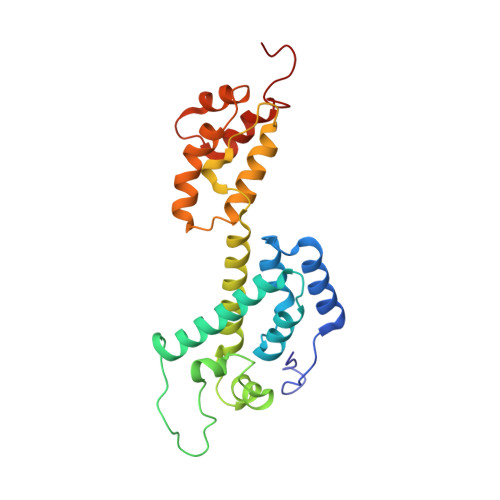
Atomic-resolution structure of HIV-1 capsid tubes by magic-angle spinning NMR.
Lu, M., Russell, R.W., Bryer, A.J., Quinn, C.M., Hou, G., Zhang, H., Schwieters, C.D., Perilla, J.R., Gronenborn, A.M., Polenova, T.(2020) Nat Struct Mol Biol 27: 863-869
- PubMed: 32901160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0489-2
- Primary Citation Related Structures:
6WAP, 6X63 - PubMed Abstract:
HIV-1 capsid plays multiple key roles in viral replication, and inhibition of capsid assembly is an attractive target for therapeutic intervention. Here, we report the atomic-resolution structure of capsid protein (CA) tubes, determined by magic-angle spinning NMR and data-guided molecular dynamics simulations. Functionally important regions, including the NTD β-hairpin, the cyclophilin A-binding loop, residues in the hexamer central pore, and the NTD-CTD linker region, are well defined. The structure of individual CA chains, their arrangement in the pseudo-hexameric units of the tube and the inter-hexamer interfaces are consistent with those in intact capsids and substantially different from the organization in crystal structures, which feature flat hexamers. The inherent curvature in the CA tubes is controlled by conformational variability of residues in the linker region and of dimer and trimer interfaces. The present structure reveals atomic-level detail in capsid architecture and provides important guidance for the design of novel capsid inhibitors.
- Department of Chemistry and Biochemistry, University of Delaware, Newark, DE, USA.
Organizational Affiliation: