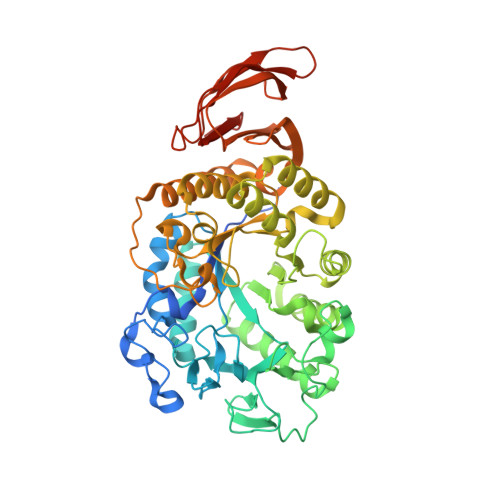
Discovery of a novel group of three-domain thermophilic cyclomaltodextrin glucanotransferases: structural and functional implications.
Centeno-Leija, S., Lopez-Munguia, A., Cardenas-Conejo, Y., Mancilla-Margalli, N.A., Velazquez-Cruz, B., Magana-Cuevas, E., Guerra-Borrego, Y., Zatarain-Palacios, R., Marin-Tovar, Y., Gomez-Manzo, S., Marcial-Quino, J., Hernandez-Ochoa, B., Osuna-Castro, J.A., Rudino-Pinera, E., Serrano-Posada, H.To be published.