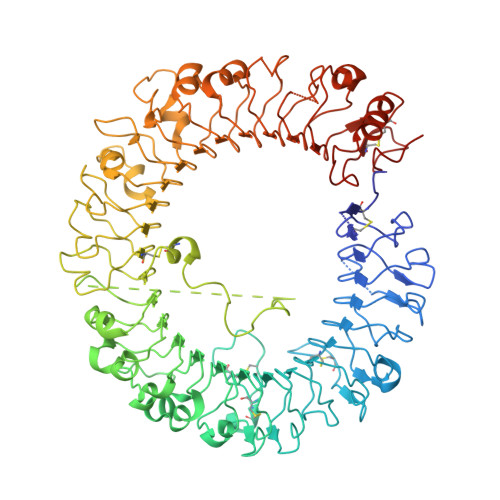
Discovery of GS-9688 (Selgantolimod) as a Potent and Selective Oral Toll-Like Receptor 8 Agonist for the Treatment of Chronic Hepatitis B.
Mackman, R.L., Mish, M., Chin, G., Perry, J.K., Appleby, T., Aktoudianakis, V., Metobo, S., Pyun, P., Niu, C., Daffis, S., Yu, H., Zheng, J., Villasenor, A.G., Zablocki, J., Chamberlain, J., Jin, H., Lee, G., Suekawa-Pirrone, K., Santos, R., Delaney 4th, W.E., Fletcher, S.P.(2020) J Med Chem 63: 10188-10203
- PubMed: 32407112 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.0c00100
- Primary Citation Related Structures:
6WML - PubMed Abstract:
Toll-like receptor 8 (TLR8) recognizes pathogen-derived single-stranded RNA fragments to trigger innate and adaptive immune responses. Chronic hepatitis B (CHB) is associated with a dysfunctional immune response, and therefore a selective TLR8 agonist may be an effective treatment option. Structure-based optimization of a dual TLR7/8 agonist led to the identification of the selective TLR8 clinical candidate ( R )-2-((2-amino-7-fluoropyrido[3,2- d ]pyrimidin-4-yl)amino)-2-methylhexan-1-ol (GS-9688, ( R )- 7 ). Potent TLR8 agonism (IL-12p40 EC 50 = 220 nM) and >100-fold TLR7 selectivity (IFN-α EC 50 > 50 μM) was observed in human peripheral blood mononuclear cells (PBMCs). The TLR8-ectodomain:( R )- 7 complex confirmed TLR8 binding and a direct ligand interaction with TLR8 residue Asp545. Oral ( R )- 7 had good absorption and high first pass clearance in preclinical species. A reduction in viral markers was observed in HBV-infected primary human hepatocytes treated with media from PBMCs stimulated with ( R )- 7 , supporting the clinical development of ( R )- 7 for the treatment of CHB.
- Gilead Sciences, Inc., 333 Lakeside Drive, Foster City, California 94404, United States.
Organizational Affiliation: