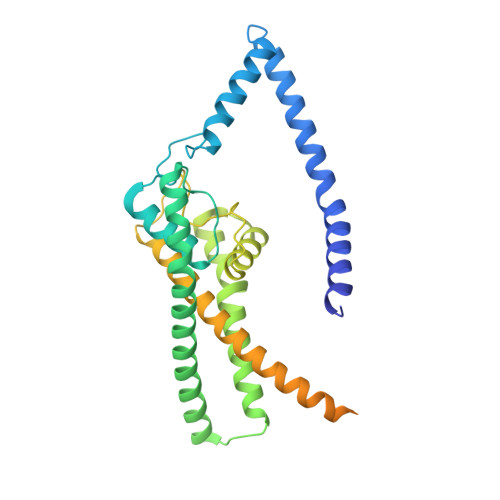
Structural basis for pH gating of the two-pore domain K + channel TASK2.
Li, B., Rietmeijer, R.A., Brohawn, S.G.(2020) Nature 586: 457-462
- PubMed: 32999458 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-020-2770-2
- Primary Citation Related Structures:
6WLV, 6WM0 - PubMed Abstract:
TASK2 (also known as KCNK5) channels generate pH-gated leak-type K + currents to control cellular electrical excitability 1-3 . TASK2 is involved in the regulation of breathing by chemosensory neurons of the retrotrapezoid nucleus in the brainstem 4-6 and pH homeostasis by kidney proximal tubule cells 7,8 . These roles depend on channel activation by intracellular and extracellular alkalization 3,8,9 , but the mechanistic basis for TASK2 gating by pH is unknown. Here we present cryo-electron microscopy structures of Mus musculus TASK2 in lipid nanodiscs in open and closed conformations. We identify two gates, distinct from previously observed K + channel gates, controlled by stimuli on either side of the membrane. Intracellular gating involves lysine protonation on inner helices and the formation of a protein seal between the cytoplasm and the channel. Extracellular gating involves arginine protonation on the channel surface and correlated conformational changes that displace the K + -selectivity filter to render it nonconductive. These results explain how internal and external protons control intracellular and selectivity filter gates to modulate TASK2 activity.
- Department of Molecular and Cell Biology, University of California Berkeley, Berkeley, CA, USA.
Organizational Affiliation: