Convergent Evolution in Breadth of Two VH6-1-Encoded Influenza Antibody Clonotypes from a Single Donor.
Wu, N.C., Andrews, S.F., Raab, J.E., O'Connell, S., Schramm, C.A., Ding, X., Chambers, M.J., Leung, K., Wang, L., Zhang, Y., Mascola, J.R., Douek, D.C., Ledgerwood, J.E., McDermott, A.B., Wilson, I.A.(2020) Cell Host Microbe 28: 434
- PubMed: 32619441 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.chom.2020.06.003
- Primary Citation Related Structures:
6WIY, 6WIZ, 6WJ0, 6WJ1 - PubMed Abstract:
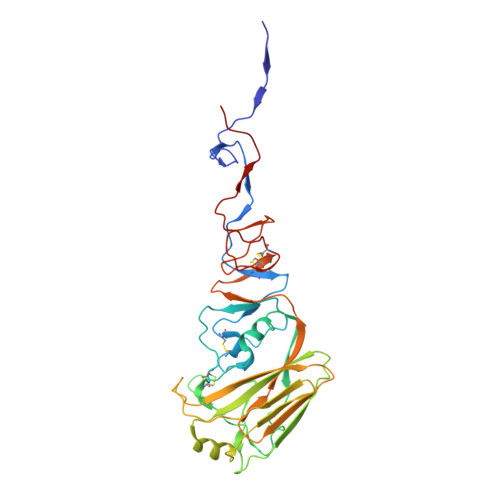
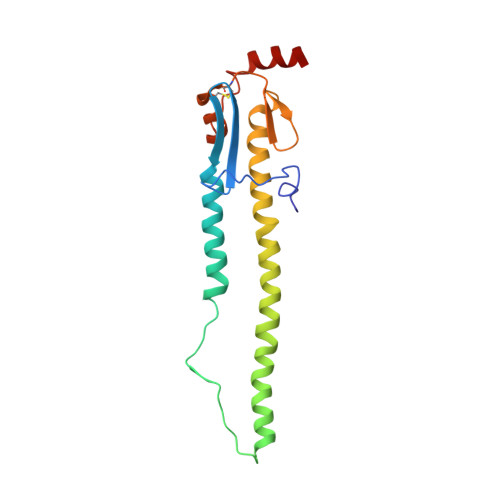
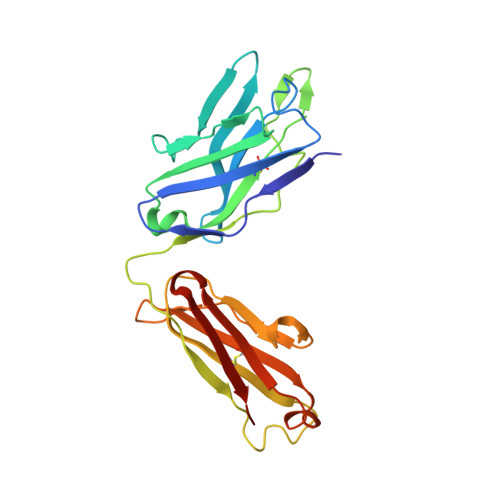
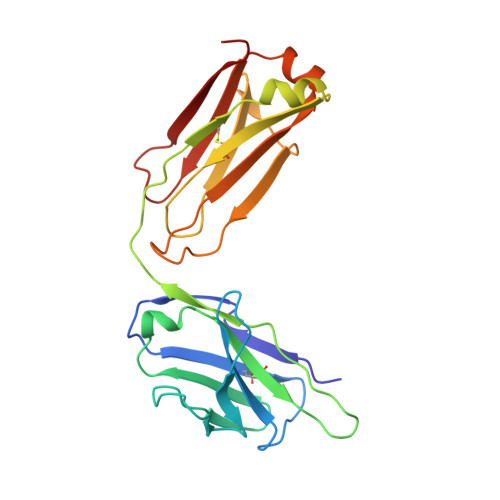
Understanding how broadly neutralizing antibodies (bnAbs) to influenza hemagglutinin (HA) naturally develop in humans is critical to the design of universal influenza vaccines. Several classes of bnAbs directed to the conserved HA stem were found in multiple individuals, including one encoded by heavy-chain variable domain V H 6-1. We describe two genetically similar V H 6-1 bnAb clonotypes from the same individual that exhibit different developmental paths toward broad neutralization activity. One clonotype evolved from a germline precursor recognizing influenza group 1 subtypes to gain breadth to group 2 subtypes. The other clonotype recognized group 2 subtypes and developed binding to group 1 subtypes through somatic hypermutation. Crystal structures reveal that the specificity differences are primarily mediated by complementarity-determining region H3 (CDR H3). Thus, while V H 6-1 provides a framework for development of HA stem-directed bnAbs, sequence differences in CDR H3 junctional regions during VDJ recombination can alter reactivity and evolutionary pathways toward increased breadth.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, San Diego, CA 92037, USA.
Organizational Affiliation: