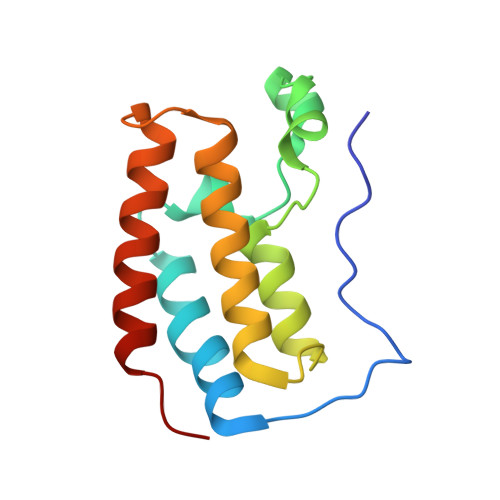
Selective N-Terminal BET Bromodomain Inhibitors by Targeting Non-Conserved Residues and Structured Water Displacement*.
Cui, H., Divakaran, A., Pandey, A.K., Johnson, J.A., Zahid, H., Hoell, Z.J., Ellingson, M.O., Shi, K., Aihara, H., Harki, D.A., Pomerantz, W.C.K.(2021) Angew Chem Int Ed Engl 60: 1220-1226
- PubMed: 32975004 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202008625
- Primary Citation Related Structures:
6WGX - PubMed Abstract:
Bromodomain and extra-terminal (BET) family proteins, BRD2-4 and T, are important drug targets; however, the biological functions of each bromodomain remain ill-defined. Chemical probes that selectively inhibit a single BET bromodomain are lacking, although pan inhibitors of the first (D1), and second (D2), bromodomain are known. Here, we develop selective BET D1 inhibitors with preferred binding to BRD4 D1. In competitive inhibition assays, we show that our lead compound is 9-33 fold selective for BRD4 D1 over the other BET bromodomains. X-ray crystallography supports a role for the selectivity based on reorganization of a non-conserved lysine and displacement of an additional structured water in the BRD4 D1 binding site relative to our prior lead. Whereas pan-D1 inhibitors displace BRD4 from MYC enhancers, BRD4 D1 inhibition in MM.1S cells is insufficient for stopping Myc expression and may lead to its upregulation. Future analysis of BRD4 D1 gene regulation may shed light on differential BET bromodomain functions.
- Department of Chemistry, University of Minnesota-Twin Cities, 207 Pleasant St. SE, Minneapolis, MN, 55455, USA.
Organizational Affiliation: