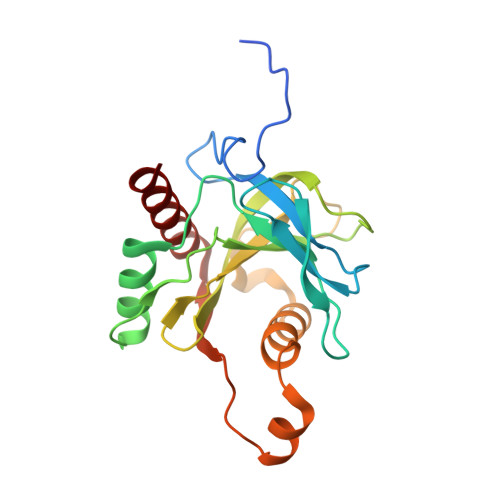
Crystal structure of an inorganic pyrophosphatase from Chlamydia trachomatis D/UW-3/Cx.
Maddy, J., Staker, B.L., Subramanian, S., Abendroth, J., Edwards, T.E., Myler, P.J., Hybiske, K., Asojo, O.A.(2022) Acta Crystallogr F Struct Biol Commun 78: 135-142
- PubMed: 35234139 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X22002138
- Primary Citation Related Structures:
6WE5 - PubMed Abstract:
Chlamydia trachomatis is the leading cause of bacterial sexually transmitted infections globally and is one of the most commonly reported infections in the United States. There is a need to develop new therapeutics due to drug resistance and the failure of current treatments to clear persistent infections. Structures of potential C. trachomatis rational drug-discovery targets, including C. trachomatis inorganic pyrophosphatase (CtPPase), have been determined by the Seattle Structural Genomics Center for Infectious Disease. Inorganic pyrophosphatase hydrolyzes inorganic pyrophosphate during metabolism. Furthermore, bacterial inorganic pyrophosphatases have shown promise for therapeutic discovery. Here, a 2.2 Å resolution X-ray structure of CtPPase is reported. The crystal structure of CtPPase reveals shared structural features that may facilitate the repurposing of inhibitors identified for bacterial inorganic pyrophosphatases as starting points for new therapeutics for C. trachomatis.
- Department of Chemistry and Biochemistry, Hampton University, 100 East Queen Street, Hampton, VA 23668, USA.
Organizational Affiliation: