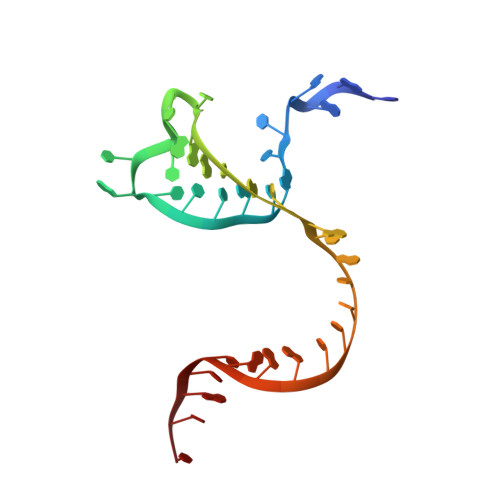
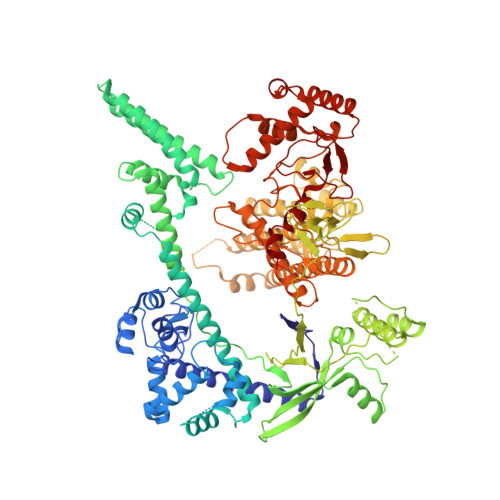
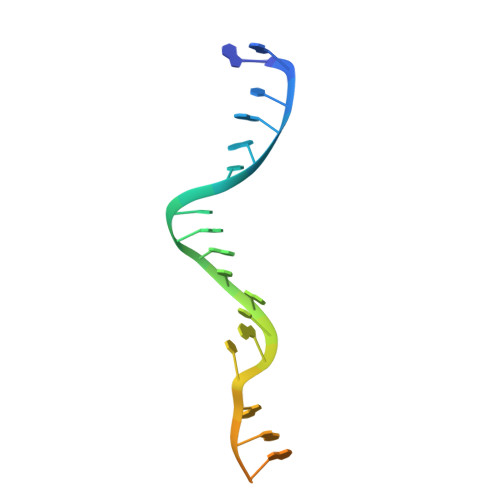
Mechanisms for target recognition and cleavage by the Cas12i RNA-guided endonuclease.
Zhang, H., Li, Z., Xiao, R., Chang, L.(2020) Nat Struct Mol Biol 27: 1069-1076
- PubMed: 32895556 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-020-0499-0
- Primary Citation Related Structures:
6W5C, 6W62, 6W64 - PubMed Abstract:
Cas12i is a recently identified type V CRISPR-Cas endonuclease that predominantly cleaves the non-target strand of a double-stranded DNA substrate. This nicking activity of Cas12i could potentially be used for genome editing with high specificity. To elucidate its mechanisms for target recognition and cleavage, we determined cryo-EM structures of Cas12i in multiple functional states. Cas12i pre-orders a seven-nucleotide seed sequence of the crRNA for target recognition and undergoes a two-step activation through crRNA-DNA hybridization. Formation of 14 base pairs activates the nickase activity, and 28-bp hybridization promotes cleavage of the target strand. The atomic structures and mechanistic insights gained should facilitate the manipulation of Cas12i for genome editing applications.
- Department of Biological Sciences, Purdue University, West Lafayette, IN, USA. zhangheng134@gmail.com.
Organizational Affiliation: