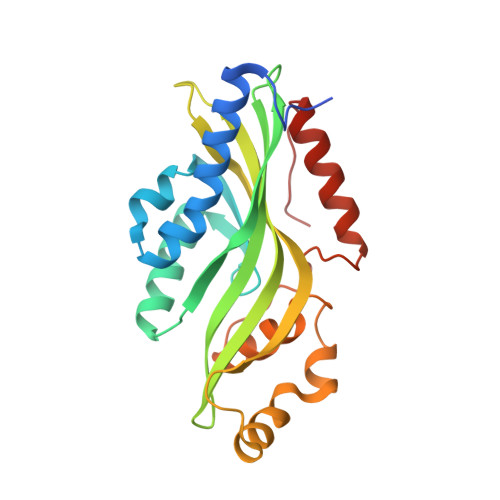
Crystal structures of non-oxidative decarboxylases reveal a new mechanism of action with a catalytic dyad and structural twists.
Zeug, M., Markovic, N., Iancu, C.V., Tripp, J., Oreb, M., Choe, J.Y.(2021) Sci Rep 11: 3056-3056
- PubMed: 33542397 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-021-82660-z
- Primary Citation Related Structures:
6W54, 7JMR, 7JMV, 7KD9 - PubMed Abstract:
Hydroxybenzoic acids, like gallic acid and protocatechuic acid, are highly abundant natural compounds. In biotechnology, they serve as critical precursors for various molecules in heterologous production pathways, but a major bottleneck is these acids' non-oxidative decarboxylation to hydroxybenzenes. Optimizing this step by pathway and enzyme engineering is tedious, partly because of the complicating cofactor dependencies of the commonly used prFMN-dependent decarboxylases. Here, we report the crystal structures (1.5-1.9 Å) of two homologous fungal decarboxylases, AGDC1 from Arxula adenivorans, and PPP2 from Madurella mycetomatis. Remarkably, both decarboxylases are cofactor independent and are superior to prFMN-dependent decarboxylases when heterologously expressed in Saccharomyces cerevisiae. The organization of their active site, together with mutational studies, suggests a novel decarboxylation mechanism that combines acid-base catalysis and transition state stabilization. Both enzymes are trimers, with a central potassium binding site. In each monomer, potassium introduces a local twist in a β-sheet close to the active site, which primes the critical H86-D40 dyad for catalysis. A conserved pair of tryptophans, W35 and W61, acts like a clamp that destabilizes the substrate by twisting its carboxyl group relative to the phenol moiety. These findings reveal AGDC1 and PPP2 as founding members of a so far overlooked group of cofactor independent decarboxylases and suggest strategies to engineer their unique chemistry for a wide variety of biotechnological applications.
- Department of Chemistry, Biochemistry, and Pharmacy, Goethe University Frankfurt, Frankfurt am Main, Germany.
Organizational Affiliation: