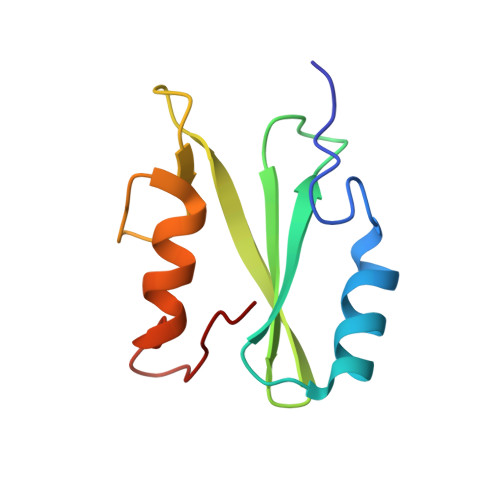
The dynamics of free and phosphopeptide-bound Grb2-SH2 reveals two dynamically independent subdomains and an encounter complex with fuzzy interactions.
Sanches, K., Caruso, I.P., Almeida, F.C.L., Melo, F.A.(2020) Sci Rep 10: 13040-13040
- PubMed: 32747626 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-70034-w
- Primary Citation Related Structures:
6VK2 - PubMed Abstract:
The growth factor receptor-bound protein 2 (Grb2) is a key factor in the regulation of cell survival, proliferation, differentiation, and metabolism. In its structure, the central Src homology 2 (SH2) domain is flanked by two Src homology 3 (SH3). SH2 is the most important domain in the recognition of phosphotyrosines. Here, we present the first dynamical characterization of Grb2-SH2 domain in the free state and in the presence of phosphopeptide EpYINSQV at multiple timescales, which revealed valuable information to the understanding of phophotyrosine sensing mechanism. Grb2-SH2 presented two dynamically independent subdomains, subdomain I involved in pY recognition and subdomain II is the pY + 2 specificity pocket. Under semi-saturated concentrations of pY-pep we observed fuzzy interactions, which led to chemical exchange observed by NMR. This information was used to describe the encounter complex. The association with pY-pep is dynamic, involving fuzzy interactions and multiple conformations of pY-pep with negative and hydrophobic residues, creating an electrostatic-potential that drives the binding of pY-pep. The recognition face is wider than the binding site, with many residues beyond the central SH2 binding site participating in the association complex, which contribute to explain previously reported capability of Grb2 to recognize remote pY.
- Multiuser Center for Biomolecular Innovation (CMIB), Department of Physics, São Paulo State University (UNESP), São Jose do Rio Preto, São Paulo, Brazil.
Organizational Affiliation: