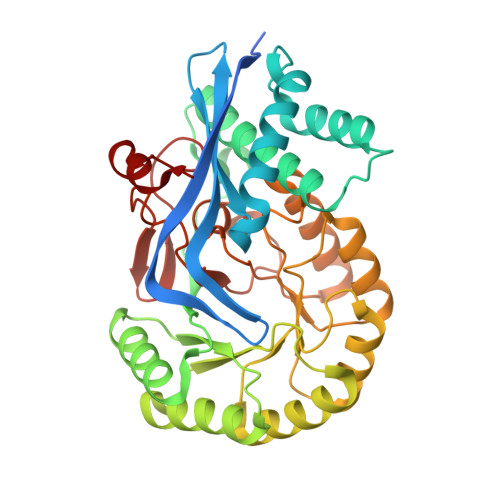
Potent Inhibition of Mandelate Racemase by Boronic Acids: Boron as a Mimic of a Carbon Acid Center.
Sharma, A.N., Grandinetti, L., Johnson, E.R., St Maurice, M., Bearne, S.L.(2020) Biochemistry 59: 3026-3037
- PubMed: 32786399 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.0c00478
- Primary Citation Related Structures:
6VIM - PubMed Abstract:
Boronic acids have been successfully employed as inhibitors of hydrolytic enzymes. Typically, an enzymatic nucleophile catalyzing hydrolysis adds to the electrophilic boron atom forming a tetrahedral species that mimics the intermediate(s)/transition state(s) for the hydrolysis reaction. We show that para -substituted phenylboronic acids (PBAs) are potent competitive inhibitors of mandelate racemase (MR), an enzyme that catalyzes a 1,1-proton transfer rather than a hydrolysis reaction. The K i value for PBA was 1.8 ± 0.1 μM, and p -Cl-PBA exhibited the most potent inhibition ( K i = 81 ± 4 nM), exceeding the binding affinity of the substrate by ∼4 orders of magnitude. Isothermal titration calorimetric studies with the wild-type, K166M, and H297N MR variants indicated that, of the two Brønsted acid-base catalysts Lys 166 and His 297, the former made the greater contribution to inhibitor binding. The X-ray crystal structure of the MR·PBA complex revealed the presence of multiple H-bonds between the boronic acid hydroxyl groups and the side chains of active site residues, as well as formation of a His 297 N ε2 -B dative bond. The dramatic upfield change in chemical shift of 27.2 ppm in the solution-phase 11 B nuclear magnetic resonance spectrum accompanying binding of PBA by MR was consistent with an sp 3 -hybridized boron, which was also supported by density-functional theory calculations. These unprecedented findings suggest that, beyond substituting boron at carbon centers participating in hydrolysis reactions, substitution of boron at the acidic carbon center of a substrate furnishes a new approach for generating inhibitors of enzymes catalyzing the deprotonation of carbon acid substrates.
- Department of Biochemistry and Molecular Biology, Dalhousie University, Halifax, NS B3H 4R2, Canada.
Organizational Affiliation: