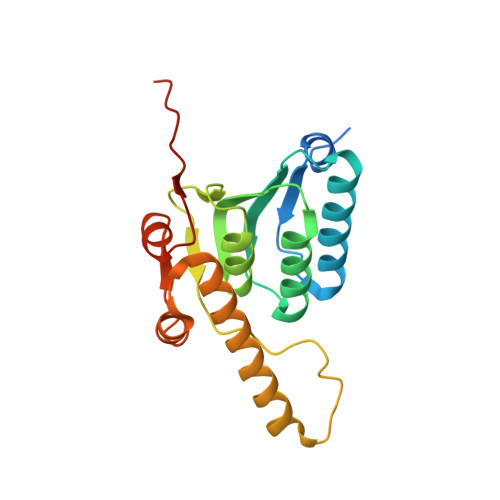
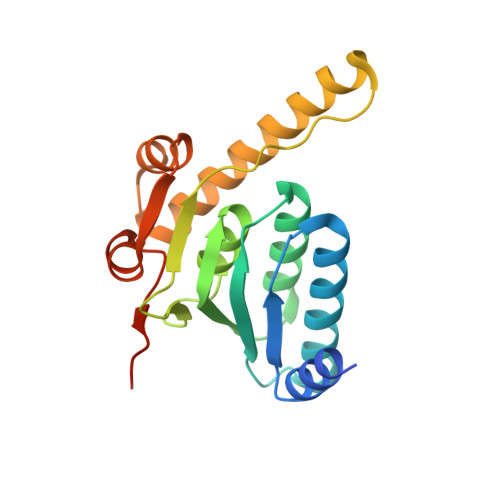
An allosteric switch regulatesMycobacterium tuberculosisClpP1P2 protease function as established by cryo-EM and methyl-TROSY NMR.
Vahidi, S., Ripstein, Z.A., Juravsky, J.B., Rennella, E., Goldberg, A.L., Mittermaier, A.K., Rubinstein, J.L., Kay, L.E.(2020) Proc Natl Acad Sci U S A 117: 5895-5906
- PubMed: 32123115 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1921630117
- Primary Citation Related Structures:
6VGK, 6VGN, 6VGQ - PubMed Abstract:
The 300-kDa ClpP1P2 protease from Mycobacterium tuberculosis collaborates with the AAA+ (ATPases associated with a variety of cellular activities) unfoldases, ClpC1 and ClpX, to degrade substrate proteins. Unlike in other bacteria, all of the components of the Clp system are essential for growth and virulence of mycobacteria, and their inhibitors show promise as antibiotics. MtClpP1P2 is unique in that it contains a pair of distinct ClpP1 and ClpP2 rings and also requires the presence of activator peptides, such as benzoyl-leucyl-leucine (Bz-LL), for function. Understanding the structural basis for this requirement has been elusive but is critical for the rational design and improvement of antituberculosis (anti-TB) therapeutics that target the Clp system. Here, we present a combined biophysical and biochemical study to explore the structure-dynamics-function relationship in MtClpP1P2. Electron cryomicroscopy (cryo-EM) structures of apo and acyldepsipeptide-bound MtClpP1P2 explain their lack of activity by showing loss of a key β-sheet in a sequence known as the handle region that is critical for the proper formation of the catalytic triad. Methyl transverse relaxation-optimized spectroscopy (TROSY)-based NMR, cryo-EM, and biochemical assays show that, on binding Bz-LL or covalent inhibitors, MtClpP1P2 undergoes a conformational change from an inactive compact state to an active extended structure that can be explained by a modified Monod-Wyman-Changeux model. Our study establishes a critical role for the handle region as an on/off switch for function and shows extensive allosteric interactions involving both intra- and interring communication that regulate MtClpP1P2 activity and that can potentially be exploited by small molecules to target M. tuberculosis .
- Department of Molecular Genetics, University of Toronto, Toronto, ON M5S 1A8, Canada.
Organizational Affiliation: