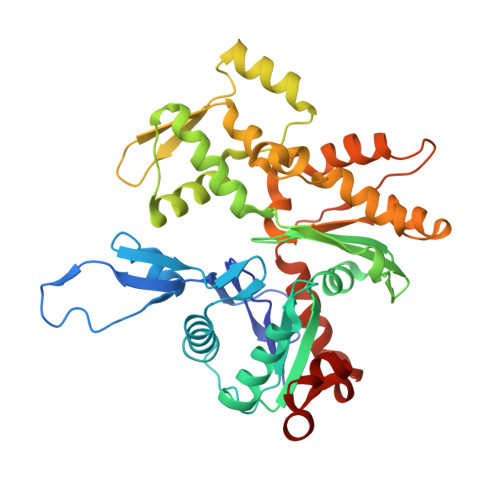
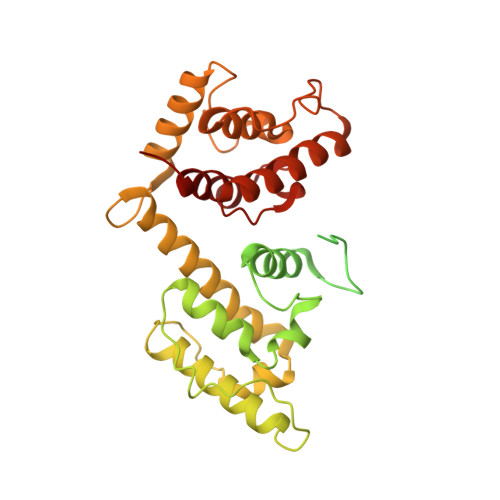
Osteogenesis imperfecta mutations in plastin 3 lead to impaired calcium regulation of actin bundling.
Schwebach, C.L., Kudryashova, E., Zheng, W., Orchard, M., Smith, H., Runyan, L.A., Egelman, E.H., Kudryashov, D.S.(2020) Bone Res 8: 21-21
- PubMed: 32509377 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41413-020-0095-2
- Primary Citation Related Structures:
6VEC - PubMed Abstract:
Mutations in actin-bundling protein plastin 3 (PLS3) emerged as a cause of congenital osteoporosis, but neither the role of PLS3 in bone development nor the mechanisms underlying PLS3-dependent osteoporosis are understood. Of the over 20 identified osteoporosis-linked PLS3 mutations, we investigated all five that are expected to produce full-length protein. One of the mutations distorted an actin-binding loop in the second actin-binding domain of PLS3 and abolished F-actin bundling as revealed by cryo-EM reconstruction and protein interaction assays. Surprisingly, the remaining four mutants fully retained F-actin bundling ability. However, they displayed defects in Ca 2+ sensitivity: two of the mutants lost the ability to be inhibited by Ca 2+ , while the other two became hypersensitive to Ca 2+ . Each group of the mutants with similar biochemical properties showed highly characteristic cellular behavior. Wild-type PLS3 was distributed between lamellipodia and focal adhesions. In striking contrast, the Ca 2+ -hyposensitive mutants were not found at the leading edge but localized exclusively at focal adhesions/stress fibers, which displayed reinforced morphology. Consistently, the Ca 2+ -hypersensitive PLS3 mutants were restricted to lamellipodia, while chelation of Ca 2+ caused their redistribution to focal adhesions. Finally, the bundling-deficient mutant failed to co-localize with any F-actin structures in cells despite a preserved F-actin binding through a non-mutation-bearing actin-binding domain. Our findings revealed that severe osteoporosis can be caused by a mutational disruption of the Ca 2+ -controlled PLS3's cycling between adhesion complexes and the leading edge. Integration of the structural, biochemical, and cell biology insights enabled us to propose a molecular mechanism of plastin activity regulation by Ca 2+ .
- Department of Chemistry and Biochemistry, The Ohio State University, Columbus, OH 43210 USA.
Organizational Affiliation: