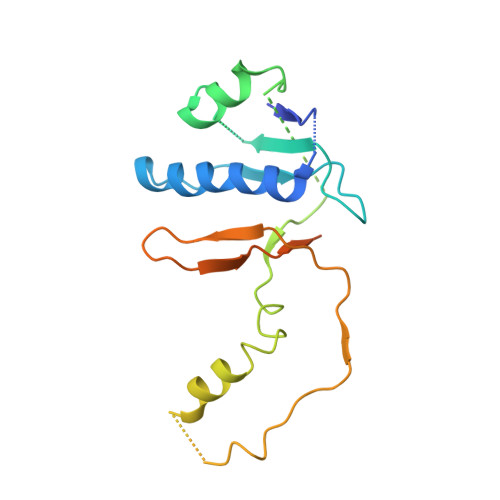
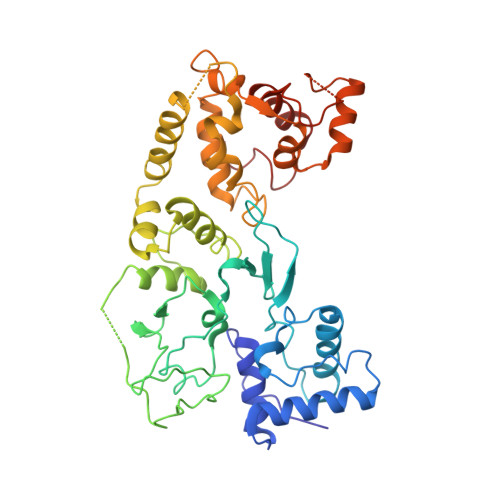
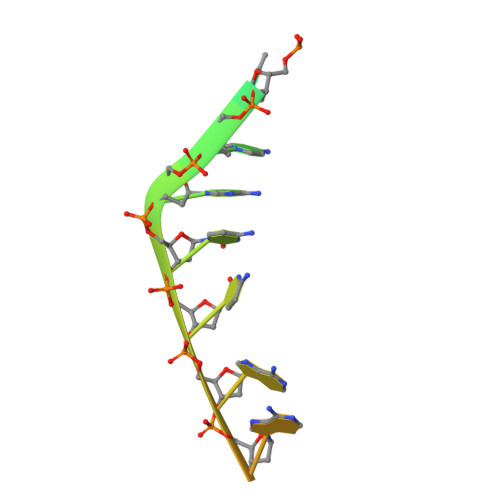
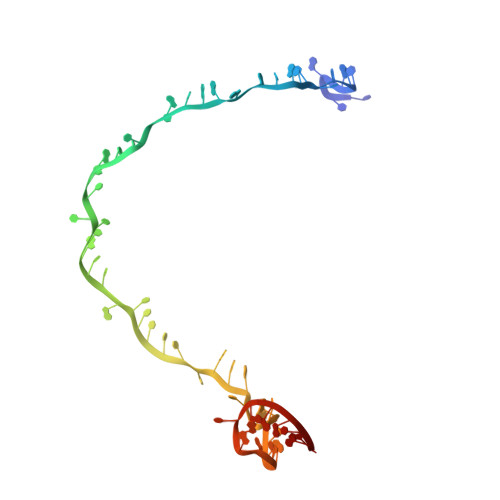
Structure-function insights into the initial step of DNA integration by a CRISPR-Cas-Transposon complex.
Jia, N., Xie, W., de la Cruz, M.J., Eng, E.T., Patel, D.J.(2020) Cell Res 30: 182-184
- PubMed: 31925391 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-019-0272-2
- Primary Citation Related Structures:
6V9P, 6V9Q, 6VBW - Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY, 10065, USA. jian@mskcc.org.
Organizational Affiliation: