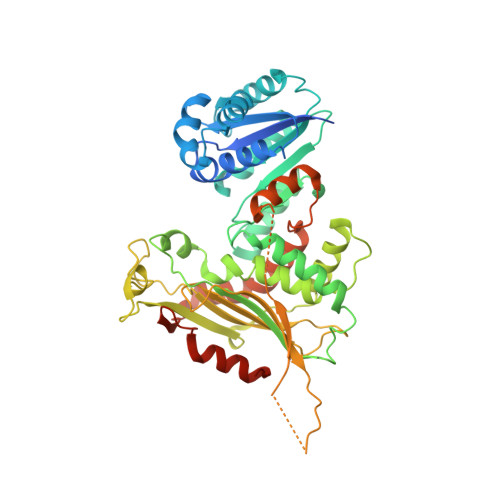
Long-range structural defects by pathogenic mutations in most severe glucose-6-phosphate dehydrogenase deficiency.
Horikoshi, N., Hwang, S., Gati, C., Matsui, T., Castillo-Orellana, C., Raub, A.G., Garcia, A.A., Jabbarpour, F., Batyuk, A., Broweleit, J., Xiang, X., Chiang, A., Broweleit, R., Vohringer-Martinez, E., Mochly-Rosen, D., Wakatsuki, S.(2021) Proc Natl Acad Sci U S A 118
- PubMed: 33468660 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2022790118
- Primary Citation Related Structures:
6VA0, 6VA7, 6VA8, 6VA9, 6VAQ - PubMed Abstract:
Glucose-6-phosphate dehydrogenase (G6PD) deficiency is the most common blood disorder, presenting multiple symptoms, including hemolytic anemia. It affects 400 million people worldwide, with more than 160 single mutations reported in G6PD. The most severe mutations (about 70) are classified as class I, leading to more than 90% loss of activity of the wild-type G6PD. The crystal structure of G6PD reveals these mutations are located away from the active site, concentrating around the noncatalytic NADP + -binding site and the dimer interface. However, the molecular mechanisms of class I mutant dysfunction have remained elusive, hindering the development of efficient therapies. To resolve this, we performed integral structural characterization of five G6PD mutants, including four class I mutants, associated with the noncatalytic NADP + and dimerization, using crystallography, small-angle X-ray scattering (SAXS), cryogenic electron microscopy (cryo-EM), and biophysical analyses. Comparisons with the structure and properties of the wild-type enzyme, together with molecular dynamics simulations, bring forward a universal mechanism for this severe G6PD deficiency due to the class I mutations. We highlight the role of the noncatalytic NADP + -binding site that is crucial for stabilization and ordering two β-strands in the dimer interface, which together communicate these distant structural aberrations to the active site through a network of additional interactions. This understanding elucidates potential paths for drug development targeting G6PD deficiency.
- Life Science Center for Survival Dynamics, University of Tsukuba, Ibaraki 305-8577, Japan.
Organizational Affiliation: