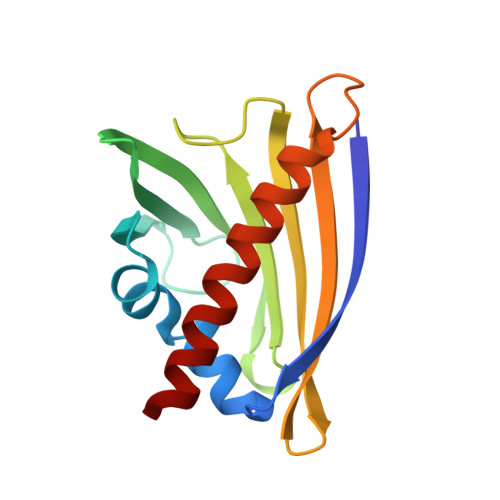
Structural analysis of 8-anilino-1-naphthalene sulfonate (ANS) binding to the PR-10 allergen Ara h 8.
O'Malley, A., Offermann, L.R., Khatri, K., Linn, C., Pote, S., McBride, J.K., Perdue, M.L., Hurlburt, B.K., Maleki, S.J., Mias, G.I., Chruszcz, M.(2025) Biochem Biophys Res Commun 793: 153013-153013
- PubMed: 41274249 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2025.153013
- Primary Citation Related Structures:
6AWR, 6V8H, 6V8J, 6V8M, 6V8S - PubMed Abstract:
We previously determined crystal structures of peanut allergen Ara h 8.0101 in the apo form as well as in complex with model ligands. These structures illustrated the varied ligand binding capabilities of PR-10s and Ara h 8's structural similarity to the major birch allergen Bet v 1. Here, we expanded on those structural studies with structures of Ara h 8.0101 and Ara h 8.0201 in complex with 8-anilino-1-naphthalene sulfonate (ANS), as well as the apo form of Ara h 8.0201. Structural studies revealed that both proteins may bind more than one ANS molecule. We also examined the impact of ANS on the ligand binding cavities of Ara h 8.0101 and Ara h 8.0201 with fluorescence assays and compared the results to prototypic PR-10 Bet v 1.0101. Moreover, as ANS is often used in fluorescence-based ligand binding assays, we analyzed structures from the PDB and provided a summary on experimentally determined ANS binding sites. These analyses show that ANS is useful for investigation of ligand binding sites, but it may also participate in non-specific reactions on nonpolar surfaces of proteins.
- Department of Biochemistry and Molecular Biology, Michigan State University, East Lansing, MI, 48824, USA.
Organizational Affiliation: