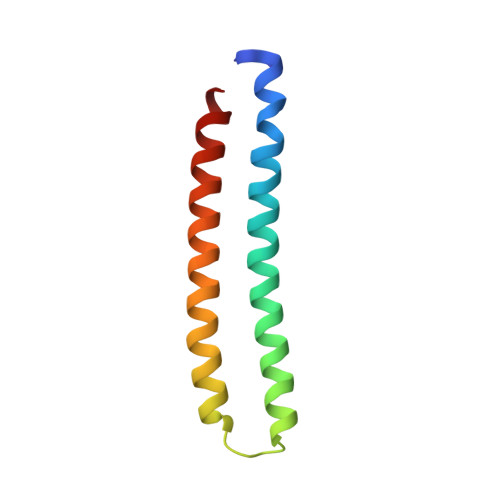
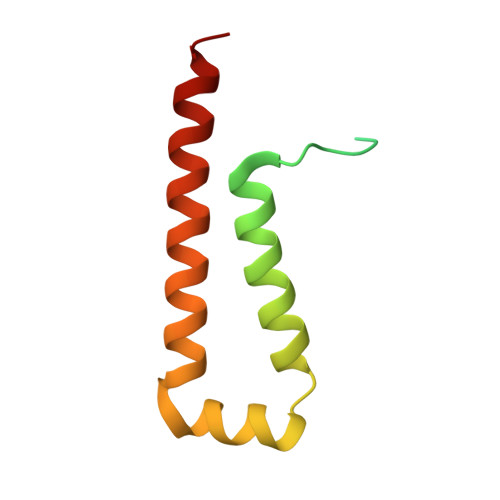
Crystal structure of a proteolytically cleaved, amino terminal domain of apolipoprotein E3.
McPherson, A.(2020) Biochem Biophys Res Commun
- PubMed: 32081433 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.01.117
- Primary Citation Related Structures:
6V7M - PubMed Abstract:
A sample of Apolipoprotein E3 used in the original structure determination by X-ray crystallography (PDB code 1NFN) was crystallized under different conditions and its structure determined by molecular replacement at 298° K. The original model (1NFN) began at amino acid 23 and ended at amino acid 164, but the amino acid segment 81 through 91 (a loop between helices) was not visible in the electron density and presumed disordered. The model reported here is essentially identical to 1NFN, but now includes amino acids 18 through 22 at the amino terminus, 165 at the carboxy terminus and includes as well the segment 83 through 91. Leu 82 is not visible, but the separation between Gln 81 and Thr 83 is more than 10 Å, thereby indicating a proteolytic cleavage occurred between those two residues.
- Dept. Molecular Biology and Biochemistry, University of California, Irvine, 3205 McGaugh Hall Irvine, CA, 92697, USA. Electronic address: amcphers@uci.edu.
Organizational Affiliation: