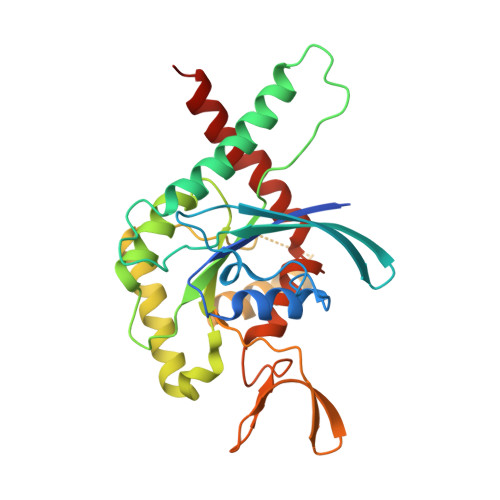
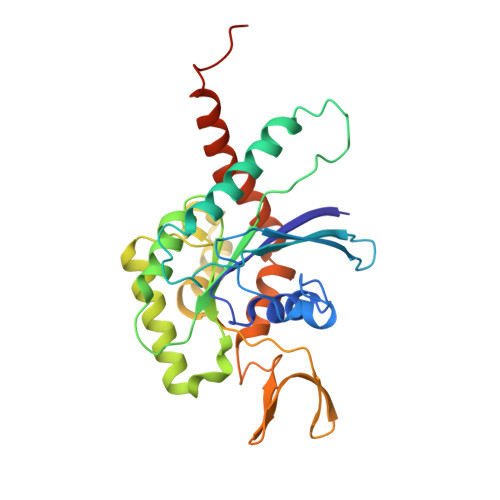
Molecular Recognition at Septin Interfaces: The Switches Hold the Key.
Rosa, H.V.D., Leonardo, D.A., Brognara, G., Brandao-Neto, J., D'Muniz Pereira, H., Araujo, A.P.U., Garratt, R.C.(2020) J Mol Biology 432: 5784-5801
- PubMed: 32910969 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2020.09.001
- Primary Citation Related Structures:
6UPA, 6UPQ, 6UPR, 6UQQ - PubMed Abstract:
The assembly of a septin filament requires that homologous monomers must distinguish between one another in establishing appropriate interfaces with their neighbors. To understand this phenomenon at the molecular level, we present the first four crystal structures of heterodimeric septin complexes. We describe in detail the two distinct types of G-interface present within the octameric particles, which must polymerize to form filaments. These are formed between SEPT2 and SEPT6 and between SEPT7 and SEPT3, and their description permits an understanding of the structural basis for the selectivity necessary for correct filament assembly. By replacing SEPT6 by SEPT8 or SEPT11, it is possible to rationalize Kinoshita's postulate, which predicts the exchangeability of septins from within a subgroup. Switches I and II, which in classical small GTPases provide a mechanism for nucleotide-dependent conformational change, have been repurposed in septins to play a fundamental role in molecular recognition. Specifically, it is switch I which holds the key to discriminating between the two different G-interfaces. Moreover, residues which are characteristic for a given subgroup play subtle, but pivotal, roles in guaranteeing that the correct interfaces are formed.
- Instituto de Física de São Carlos, Universidade de São Paulo, Avenida João Dagnone 1100, São Carlos, SP 13563-723, Brazil.
Organizational Affiliation: