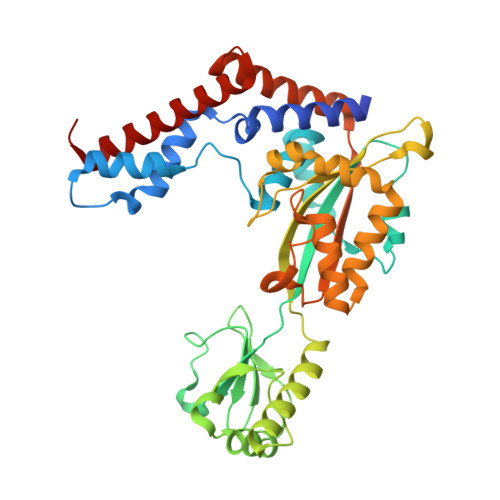
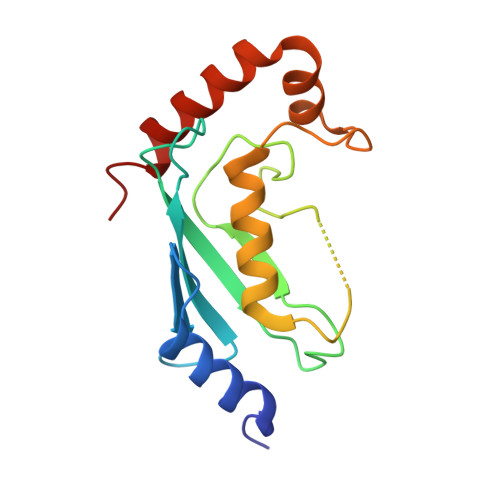
Legionella effector MavC targets the Ube2N~Ub conjugate for noncanonical ubiquitination.
Puvar, K., Iyer, S., Fu, J., Kenny, S., Negron Teron, K.I., Luo, Z.Q., Brzovic, P.S., Klevit, R.E., Das, C.(2020) Nat Commun 11: 2365-2365
- PubMed: 32398758 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-16211-x
- Primary Citation Related Structures:
6P5B, 6P5H, 6ULH, 6UMP, 6UMS - PubMed Abstract:
The bacterial effector MavC modulates the host immune response by blocking Ube2N activity employing an E1-independent ubiquitin ligation, catalyzing formation of a γ-glutamyl-ε-Lys (Gln40 Ub -Lys92 Ube2N ) isopeptide crosslink using a transglutaminase mechanism. Here we provide biochemical evidence in support of MavC targeting the activated, thioester-linked Ube2N~ubiquitin conjugate, catalyzing an intramolecular transglutamination reaction, covalently crosslinking the Ube2N and Ub subunits effectively inactivating the E2~Ub conjugate. Ubiquitin exhibits weak binding to MavC alone, but shows an increase in affinity when tethered to Ube2N in a disulfide-linked substrate that mimics the charged E2~Ub conjugate. Crystal structures of MavC in complex with the substrate mimic and crosslinked product provide insights into the reaction mechanism and underlying protein dynamics that favor transamidation over deamidation, while revealing a crucial role for the structurally unique insertion domain in substrate recognition. This work provides a structural basis of ubiquitination by transglutamination and identifies this enzyme's true physiological substrate.
- Department of Chemistry, Purdue University, West Lafayette, IN, 47907, USA.
Organizational Affiliation: