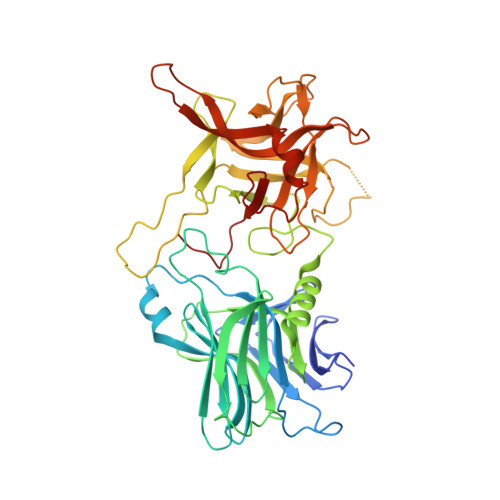
Structural Insights into Rational Design of Single-Domain Antibody-Based Antitoxins against Botulinum Neurotoxins
Lam, K., Tremblay, J.M., Vazquez-Cintron, E., Perry, K., Ondeck, C., Webb, R., McNutt, P.M., Shoemaker, C.B., Jin, R.(2020) Cell Rep 30: 2526