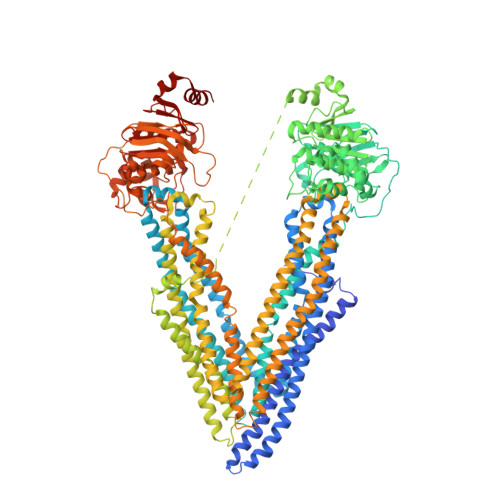
Structural definition of polyspecific compensatory ligand recognition by P-glycoprotein.
Le, C.A., Harvey, D.S., Aller, S.G.(2020) IUCrJ 7: 663-672
- PubMed: 32695413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252520005709
- Primary Citation Related Structures:
6UJN, 6UJP, 6UJR, 6UJS, 6UJT, 6UJW - PubMed Abstract:
The multidrug transporter P-glycoprotein (Pgp)/ABCB1/MDR1 plays an important role in multidrug resistance (MDR) and detoxification owing to its ability to efflux an unusually large and chemically diverse set of substrates. Previous phenylalanine-to-alanine scanning mutagenesis of Pgp revealed that nearly all mutations retained full MDR function and still permitted substrate transport. This suggests that either the loss of any single aromatic side chain did not affect the ligand-binding modes or that highly adaptive and compensatory drug recognition is an intrinsic property including ligand-binding shifts that preserve function. To explore this hypothesis, the ATPase function and crystallographic localization of five single-site mutations in which the native aromatic residue directly interacted with the environmental pollutant BDE-100, as shown in previous crystal structures, were tested. Two mutants, Y303A and Y306A, showed strong BDE-100 occupancy at the original site (site 1), but also revealed a novel site 2 located on the opposing pseudo-symmetric half of the drug-binding pocket (DBP). Surprisingly, the F724A mutant structure had no detectable binding in site 1 but exhibited a novel site shifted 11 Å from site 1. ATPase studies revealed shifts in ATPase kinetics for the five mutants, but otherwise indicated a catalytically active transporter that was inhibited by BDE-100, similar to wild-type Pgp. These results emphasize a high degree of compensatory drug recognition in Pgp that is made possible by aromatic amino-acid side chains concentrated in the DBP. Compensatory recognition forms the underpinning of polyspecific drug transport, but also highlights the challenges associated with the design of therapeutics that evade efflux altogether.
- Department of Pharmacology and Toxicology, University of Alabama at Birmingham, Birmingham, AL 35294, USA.
Organizational Affiliation: