The structure of the Streptococcus gordonii surface protein SspB in complex with TEV peptide provides clues to the adherence of oral streptococcal adherence to salivary agglutinin
Schormann, N., Deivanayagam, C.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Surface protein adhesin | 425 | Streptococcus mutans | Mutation(s): 0 Gene Names: spaP | 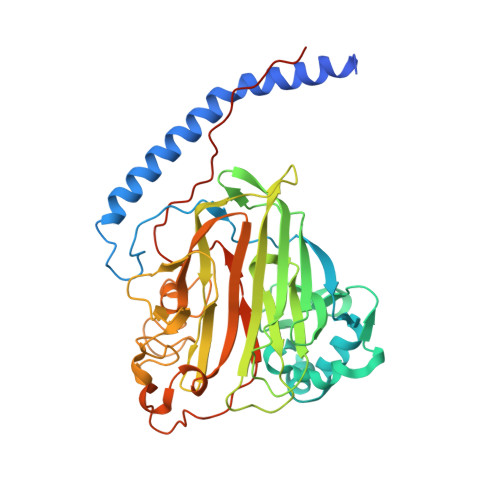 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P11657 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| SNZ (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth A], I [auth B], M [auth C], R [auth D] | N-(3,4-dihydroxyphenyl)-N'-[2-(3,4-dihydroxyphenyl)ethyl]urea C15 H16 N2 O5 XPAUTCCLOUPUGJ-UHFFFAOYSA-N |  | ||
| SO4 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | J [auth B], P [auth D] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| EDO (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | K [auth B], O [auth C], T [auth D] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| CA (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A], H [auth B], L [auth C], Q [auth D] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| NA (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | G [auth A], N [auth C], S [auth D] | SODIUM ION Na FKNQFGJONOIPTF-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 65.81 | α = 90 |
| b = 132.697 | β = 90 |
| c = 243.819 | γ = 90 |
| Software Name | Purpose |
|---|---|
| Aimless | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| DIALS | data reduction |
| PHASER | phasing |