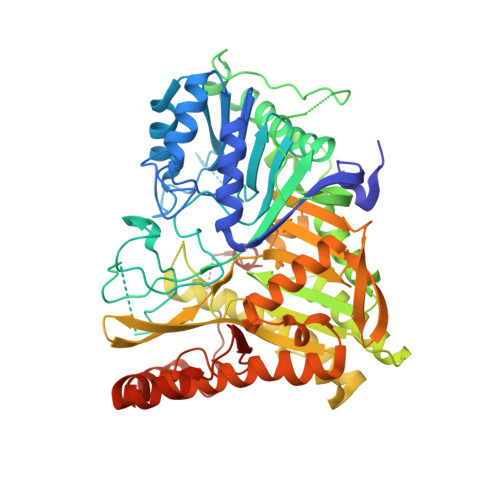
Crystal structure of human PLD1 provides insight into activation by PI(4,5)P2and RhoA.
Bowling, F.Z., Salazar, C.M., Bell, J.A., Huq, T.S., Frohman, M.A., Airola, M.V.(2020) Nat Chem Biol 16: 400-407
- PubMed: 32198492 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-020-0499-8
- Primary Citation Related Structures:
6U8Z - PubMed Abstract:
The signal transduction enzyme phospholipase D1 (PLD1) hydrolyzes phosphatidylcholine to generate the lipid second-messenger phosphatidic acid, which plays roles in disease processes such as thrombosis and cancer. PLD1 is directly and synergistically regulated by protein kinase C, Arf and Rho GTPases, and the membrane lipid phosphatidylinositol-4,5-bisphosphate (PIP 2 ). Here, we present a 1.8 Å-resolution crystal structure of the human PLD1 catalytic domain, which is characterized by a globular fold with a funnel-shaped hydrophobic cavity leading to the active site. Adjacent is a PIP 2 -binding polybasic pocket at the membrane interface that is essential for activity. The C terminus folds into and contributes part of the catalytic pocket, which harbors a phosphohistidine that mimics an intermediate stage of the catalytic cycle. Mapping of PLD1 mutations that disrupt RhoA activation identifies the RhoA-PLD1 binding interface. This structure sheds light on PLD1 regulation by lipid and protein effectors, enabling rationale inhibitor design for this well-studied therapeutic target.
- Department of Biochemistry and Cell Biology, Stony Brook University, Stony Brook, NY, USA.
Organizational Affiliation: